
DNA crosslinking damage and cancer - a tale of friend and foe
Introduction
DNA crosslinking damage occurs when crosslinking agents covalently connect two nucleotide residues from the same DNA strand (intrastrand crosslink) or from opposite strands [interstrand crosslink (ICL)]. Intrastrand crosslinks can be readily removed by the nucleotide excision repair (NER) mechanism (1). An ICL, however, constitutes an absolute block to DNA strand separation, thus interrupting essential DNA metabolic processes such as replication and transcription. Left unrepaired, ICLs can be extremely toxic especially in dividing cells, stalling DNA replication and leading to cell death. As such, ICL-forming antitumor drugs including melphalan and cisplatin are among the most widely-used chemotherapeutic agents. The first clinical application of an ICL drug, nitrogen mustard, dated back to the 1940s (2).
Earlier understanding of ICL repair mechanism was obtained primarily from studies in model systems such as Escherichia coli and yeast (3). The availability of genetic mutants and well-defined biochemical assays made it possible to establish the first ICL repair pathway, which combines NER and homologous recombination. More recently, investigations in patients with Fanconi anemia (FA) revealed an important mechanism exclusive in higher eukaryotes (vertebrates). FA patient cells are hypersensitive to ICLs as demonstrated by reduced survival rates and elevated chromosomal abnormalities. Genomic instability of FA patients is closely correlated with cancer development. Understanding of the ICL repair mechanism and their roles in cancer development and treatment is extremely important for patients with ICL repair-related diseases and for ICL-based chemotherapeutics. In this review, we describe the basic concepts of ICL repair and their implications in cancer development and treatment.
Formation of interstrand crosslinks
Formation of DNA crosslinks relies on two independently reactive groups in a single alkylating molecule. When the two reactive groups react with two bases residing on opposing DNA strands, an ICL is formed. The covalently targeted sites on DNA are usually the N7 position of guanine or the exocyclic N2-amino group of guanine from nucleotide residues from the opposite strands (4).
ICL agents exist from naturally occurring as well as synthetic sources. Naturally occurring crosslinking agents include psoralens, mitomycin C, nitrous acids, etc. Mitomycin C was originally found in fungi with antibiotic activity mediated by its DNA crosslinking ability since bacteria are easily killed by a single unrepaired ICL (5). Psoralens are natural compounds derived from plants. Psoralens target specifically the TA sequences and react with the opposing thymines to form ICLs upon photo activation by UV radiation. Because of this unique feature, psoralens are effective ectopical treatment of psoriasis (6). Certain metabolites of alcohol, cigarette, and high fat diet, such as acetaldehyde and malondialdehyde, also act as DNA interstrand as well as DNA-protein crosslinkers. Recent studies demonstrated that ICL repair-deficient mice are sensitive to aldehyde (7-9), indicating that these endogenous crosslinking agents present an internal risk of genomic instability. Another endogenous interstrand crosslinking agent is nitrous acid, which is formed in the stomach during consumption of nitrite-containing food additives.
Synthetic ICL agents consist of a broad array of bi-functional alkylators such as nitrogen mustard, carmustine, platinum compounds, and diepoxybutane. Nitrogen mustard gas was used as a chemical weapon during World War I. The observation that white blood cell counts decreased drastically from nitrogen mustard exposure led to the exploratory application of this compound in cancer therapy. Nitrogen mustard was used as a chemotherapeutic agent for lymphoma and leukemia for a period of time (10,11). Since then, many more ICL agents have been used for cancer treatment, including derivatives of nitrogen mustard such as melphalan and cyclophosphamide, and platinum-containing drugs such as cisplatin and caboplatin.
As described above, human bodies are subjected to endogenous and therapeutic ICL exposure. As an exceedingly genotoxic and cytotoxic DNA lesion, one unrepaired DNA ICL could yield lethality in monocellular organisms whereas 20 to 40 unrepaired ICLs are fatal to mammalian cells (5,12). As a result, ICL repair mechanisms are essential in maintaining genomic integrity and cell viability.
ICL repair mechanisms
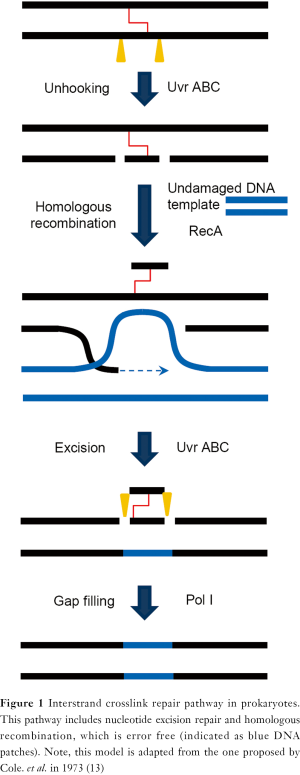
ICL repair mechanism is highly conserved in most unicellular organisms. Studies in Escherichia coli in the early 1970s demonstrated a recombination-dependent and error-free ICL repair pathway. In this model (Figure 1), NER factors initiate a strand-unhooking step by introducing dual incisions flanking the ICL lesion. Homologous recombination then fills the resulting gap by invading an undamaged chromosome. A subsequent round of NER reaction removes the remaining lesion and results in error-free repair of ICLs (13,14). This model, also called the Cole’s model (13), is supported by both genetic and biochemical evidence (15-18). A similar repair mechanism also operates in yeast (19,20). A minor ICL response pathway was shown to be recombination-independent since the recombinase RecA is not required. It was shown that polymerase Pol β is involved in a lesion bypass process leading to error-prone ICL removal (21,22).
In higher eukaryote and particularly in vertebrate cells, a complex ICL processing mechanism has evolved to facilitate ICL damage response. The importance of this pathway is reflected by the hypersensitivity of FA patient cells to crosslinking agents. Essentially, two types of mechanisms for ICL removal have been observed in eukaryotes: recombination-dependent and recombination-independent.
The recombination-dependent ICL repair pathway, alternatively termed as replication-dependent ICL repair, functions during late S or G2 phases of the cell cycle, where the ICL-damage sites are adjacent to an undamaged sister chromatid. Initiation of this repair mechanism depends on stalling of the replication fork, and the lesion removal process includes translesion synthesis, homologous recombination, and NER. In a model (Figure 2) based on studies from Xenopus laevis egg extracts (23), two replication forks converge at the ICL site and are both blocked. Unhooking of ICL takes place when structure-specific nucleases such as XPF/ERCC1 make dual incisions flanking the crosslinked site on the same DNA strand. This step releases the covalent linkage between the complementary strands. The resulting gap is filled by translesion polymerases to bypass the remaining lesion and join the downstream Okazaki fragment. The restored duplex DNA, with the crosslinked oligonucleotides attached to one strand, is further repaired by NER factors. This fully repaired sister chromatid is subsequently utilized to remove the double strand break on the sister chromatid via a classical homologous recombination pathway.
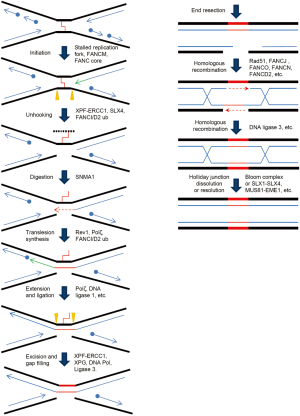
The FA pathway is involved in ICL repair from the initial recognition of stalled forks to the final step of homologous recombination (24). Two important steps seem to require the FA mechanism in the recombinational ICL repair. First, an active FA pathway is crucial for the recruitment of incision factors, as two of the potential incision factors, SLX4 and FAN1, depend on monoubiquitinated FANCD2 for their loading to ICL site via ubiquitin-binding domains (25,26). Secondly, FA pathway may tether the secondary NER incision with the ensuing homologous recombination to restore the disconnected sister chromatid (27).
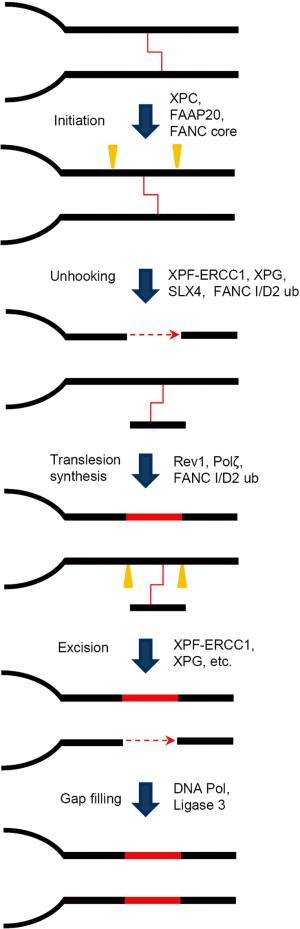
The recombination-independent ICL repair mechanism, also called mutagenic ICL repair, mainly occurs in cells during the G1 phase (Figure 3). Initiation of recombination-independent ICL repair depends on the native damage recognition proteins such as XPC and utilizes the NER dual incision to achieve the unhooking step (28,29). NER nucleases, XPF/ERCC1 and XPG provide the 5' and 3' incision, respectively. Repair synthesis of the resulting gap is aided by lesion bypass DNA polymerase ζ (30). The involvement of lesion-bypass polymerases in the recombination-independent ICL repair dictates the mutagenic nature of this pathway, which also underlines why cancer patients under ICL agent treatment are prone to chemotherapy-induced secondary malignancies (31,32).
Factors in ICL repair pathways
As described in the previous model (Figure 2), proteins involved in ICL repair include but are not limited to the 15 known FA genes (A, B, C, D1, D2, E, F, G, I, J, L, M, N, O, and P) (33-35). Essential recombination factors such as RAD51, structure-specific endonucleases such as XPF/ERCC1 and MUS81/EME1, Holliday junction processing factors, and translesion DNA polymerases are also parts of the orchestrated process during ICL repair.
FA is a rare human genetic disease characterized by pancytopenia, a broad spectrum of developmental abnormalities, and a high risk of cancer (36). Each FA subtype is associated with a distinct gene encoding a corresponding FA protein. Cells derived from FA patients exhibit high levels of chromosomal breakage and formation of radial chromosomes (37,38), indicating that ICL repair-deficient cells have high levels of genomic instability. In the classical FA pathway, FA core complex (consisting of A, G, FAAP20, C, E, F, B, L, and FAAP100) has E3 ubiquitin ligase activity with the catalytic function attributed to the RING domain-containing FANCL protein. The major function of the core complex is to execute DNA damage-induced monoubiquitination of the FANCI/D2 complex (39). The activated FANCI/D2 complex is suggested to recruit downstream effectors, including nucleases, translesion polymerases, and homologous recombination factors to repair ICLs (25,26,40). The exact role of FANCD2 monoubiquitination remains unclear.
FANCM is a DEAH domain helicase with ATP-dependent DNA translocase activity (41,42). FANCM forms a complex with FAAP24 and with the MHF1/MHF2 histone-fold complex (43,44). FANCM is important but not essential in the activation of the FA pathway (45). Biochemical studies suggest that the FANCM/FAAP24 complex stabilizes and remodels stalled DNA replication forks (44,46,47). FAAP24 complex is found to play a role in ATR-mediated checkpoint activation (48-50), whereas FANCM is shown to be involved in recombination-independent ICL repair by promoting PCNA ubiquitination thus facilitating the recruitment of NER incision factors to the ICL sites (45).
The FA gene group consisting of FANCD1 (BRCA2), FANCN, FANCJ, and FANCO are also previously established recombination factors that connect FA with breast/ovarian cancer susceptibility. Mutations of both alleles of these genes lead to a corresponding subset of FA, whereas mutation of one allele causes breast cancer predisposition. The recombination factors are likely to act downstream of ICL processing, especially when DNA double strand break forms (51,52). FANCO, also named RAD51C, is a paralogs of RAD51 (35,53). RAD51C forms complexes with RAD51B, RAD51D, XRCC2, and XRCC3 (54,55). One of the major roles of these paralogs is the recruitment and regulation of the recombinase RAD51 onto single-stranded DNA (54). Cells deficient in any of the RAD51 paralogs are sensitive to ICLs and double strand breaks because of the resulting deficit in homologous recombination (56).
Three heterodimeric structure-specific endonucleases involved in ICL repair are XPF/ERCC1, SLX1/SLX4, and MUS81/EME1. SLX4 is found to be mutated in the FANCP complementation group (34,57,58). SLX4 and SLX1 form a heterodimeric nuclease that functions as a Holliday junction resolvase in ICL repair (59-62). During ICL repair, SLX4 also serves as a scaffold protein to assemble a multi-activity nuclease complex involving XPF/ERCC1 and MUS81/EME1. XPF/ERCC1 functions in both NER and ICL repair (63,64). The NER activity of XPF/ERCC1 is SLX4 independent, which was supported by studies demonstrating that FANCP patient cells were not sensitive to UV radiation (57,65). A recent study also demonstrated that SLX4-dependent XPF/ERCC1 activity in replication-dependent ICL repair is to complete the unhooking during ICL repair (65). The unhooked oligonucleotides might be digested by another nuclease, SNM1A (66), which provide a promising alternative to the more difficult lesion bypass synthesis step.
In addition to the heterodimeric nucleases, the newly discovered nuclease FAN1 plays a moderate role in ICL repair. FAN1 is recruited to the ICL sites by the ubiquitinated FANCD2 via an ubiquitin-binding zinc finger domain in FAN1 (25). The other important domain of FAN1, the virus-type replication-repair nuclease domain, displays 5'-3' exonuclease activity and structure-specific 5'-flap endonuclease activity (25,67-69). These activities enable FAN1 to excise the exposed DNA ends as well as the stalled DNA replication structures.
Translesion DNA polymerases are important components of ICL repair. Normal replicative DNA polymerases are usually blocked ahead of the ICL site. Studies in Xenopus laevis egg extracts demonstrated that the translesion polymerases including Y-family polymerase Rev1 and the B-family polymerase Pol ζ (Rev3/Rev7) have essential roles in the complete removal of ICLs. In this models, the ICL repair intermediate utilizes replisome remodeling machinery to extend the stalled DNA strand to one base ahead of the ICL site (23). Upon unhooking, the deoxycytidyl transferase of Rev1 inserts a cytosine into the position across the ICL lesion on the complementary strand (70,71), followed by Pol ζ to extend the unpaired strand.
Interstrand crosslinking damage and development of cancer
Unrepaired or misrepaired DNA ICLs are major sources of genomic instability. ICL-inducing agents are known to be potent carcinogens. Nitrogen mustard gas exposure, aside from acute impacts, is known to cause high incidence of cancer especially leukemia (72). The onset of acute myeloid leukemia is much higher in cancer patients treated with ICL agents than those without (31,32). Clinical observations clearly indicate that administration of ICL drug has accumulative effect and the leukemogenicity of these agents is dose-dependent. However, the risk of developing leukemia is justifiable considering the benefit of chemotherapy for advanced primary malignancies.
Studies in animal models in recent years suggest that acetaldehyde, an endogenous metabolic product from alcohol, was carcinogenic (73). For example, rats exposed to acetaldehyde vapor had increased risk of squamous cell carcinoma in the respiratory epithelium, including nasal and laryngeal carcinomas. Epidemiological studies also support this notion as alcohol consumption has been linked to increased tumor incidents (74,75). More recently, mice defective in both FA pathways and Aldehyde Dehydrogenase 2 are found to be embryonic lethal (8,9). The types of DNA damage, as a result of accumulating aldehyde, may be both DNA crosslinks and DNA-protein crosslinks.
Mutation of ICL repair genes resulting failure in ICL repair, mimics ICL agent exposure. FA patients in particular exhibit high risk of hematopoietic malignancies, including myelodysplastic syndrome and acute myeloid leukemia, which account for 52% of tumors in FA patients by the age of 40 (76). The risk of all cancers, solid tumors, and acute myeloid leukemia is 50-fold, 48-fold, and 800 fold higher, respectively, in FA patients than in the general population (77). Of the solid tumors diagnosed in FA patients, head and neck squamous cell carcinoma is the most common (700-fold increase in risk) followed by esophageal cancer and gynecological cancers. The FA patients with FANCN or FANCD1 mutations usually exhibit early onset of cancer during childhood, contributing to the early mortality (78-81). FA patients defective in the homologous recombination process, (FANCD1, FANCN, and FANCJ) are more prone to breast cancer (82-84), whereas FANCO mutation carriers are more susceptible to ovarian cancer (53,85). The varying spectrum of cancer susceptibility and time of tumor onsets from different groups of FA patients underline distinct functions of FA genes during ICL removal.
Two potential mechanisms may also account for ICL-related cancers in FA patients. First, unrepaired DNA ICLs is a strong apoptosis inducer in hematopoietic cells (86), the resulting selection pressure and an increased requirement of clonogenicity for the remaining hematopoietic stem cells proved to be a key factor in the heightened risk of hematopoietic dysplasia and cancer in FA patients (87). Second, FA patients are known to have an increased risk to human papilloma virus or other oncogenic viral infection, which increases the risk of head and neck squamous cell carcinoma and other solid tumors (88). Current model suggests that the typical extension of G2 phase in FA cells (from unrepaired ICLs) increases the susceptibility to human papilloma virus infection. About 85% of head and neck squamous cell carcinoma in FA patients are positive for human papilloma virus (89).
ICL and cancer therapy
Because DNA ICLs are profoundly cytotoxic and are especially effective in killing dividing cells, ICL-inducing agents are widely used for cancer treatment, especially for solid tumors. These agents include but are not limited to nitrogen mustards, platinums, mitomycin C, and psoralens.
Clinical use of nitrogen mustards dates back about 70 years. Two of these agents, cyclophosphamide and melphalan, are still administrated as front line treatments for many forms of leukemia and myeloma. Cyclophosphamide, also known as cytophosphane with trade names including Cytoxan, Endoxan, Neosar, Procytox, and Revimmune, is routinely used in treatment of lymphoma and some types of leukemia (90) and in phase 3 clinical trials for treatment of node-positive breast cancer (91). The adverse effects of cyclophosphamide administrated at high doses include neutropenia and acute myeloid leukemia, which is the major limiting factor (92). The original clinical use of melphalan, also known as L-Phenylalanine Mustard and the trade name Alkeran, was melanoma treatment. Later, it proved to be more effective in treating myeloma (93,94). Currently, melphalan is a standard regimen for multiple myeloma (95). It is occasionally used in ovarian cancer and melanoma, though. The primary side effect of this agent is bone marrow suppression.
Other nitrogen mustard derivatives used in chemotherapy include chlorambucil (Leukeran) and ifosfamide (Ifex). Chlorambucil is primarily used for chronic lymphocytic leukemia, but replaced with Fludarabine in pediatric patients for the management of neural and bone marrow toxicity (96). Ifosfamide is used for various cancers, including testicular, lung, and breast cancer with similar side effects. Interestingly, the ifosfamide metabolite chloroacetaldehyde has chemical properties similar to those of acetaldehyde and chloral hydrate, which may explain its encephalopathy complications (97).
Platinum compounds are another class of DNA ICL-inducing agent. Cisplatin is widely used in the treatment of various solid tumors including lung cancer, ovarian cancer, lymphoma, and testicular cancer. The cure rate for testicular cancer increased from 10% to 81% with the use of adjuvant therapy with cisplatin (98). The main side effects of treatment with cisplatin are nephrotoxicity, neurotoxicity, and bone marrow suppression. Carboplatin is a second-generation platinum drug with less severe side effects in the kidney. Platinum agents developed more recently include oxaliplatin, satraplatin, picoplatin, nedaplatin, and triplatin.
In addition to the nitrogen mustard- and platinum-based drugs, mitomycin C and psoralens are also common ICL-based anticancer drugs. Mitomycin C is often used to treat esophageal, breast, and bladder cancer. The main toxic effect of intravenous mitomycin C is bone marrow suppression. Psoralens have an advantage over other ICL-based chemotherapy in the treatment of skin cancer in that psoralen-mediated ICLs are formed upon UV photo activation. Psoralens are given topically to treat cutaneous T-cell lymphoma within a defined surface area. The most common side effect of treatment with psoralens is dermatitis, which has appeared in long-term follow-up studies (99). Furthermore, psoralen exposure-induced secondary cancer is also reported from psoriasis treatment, presumably from mutations arisen from mutagenic ICL repair (100,101).
Cancers bearing BRCA2 mutations, such as breast and ovarian cancer, respond better to ICL-based chemotherapy than do cancers without these mutations. The main reason is that cancer cells with deficient homologous recombination mechanism are deficient in the later stage of ICL repair and thus sensitive to this chemotherapy. Therefore, genetic mutations affecting ICL repair actually provide tumor selectivity for ICL-based therapy. For example, platinum-based chemotherapy has improved survival rates in patients with ovarian cancer carrying BRCA2 mutations over those in patients with sporadic ovarian cancer (102,103). However, solid tumors in FA patients are mostly treated with radiation and surgery rather than ICL agents, because somatic cells in FA patients are overly sensitive to ICL agents, even much reduced ICL exposure can be fatal.
Given the effectiveness of ICL-based chemotherapy, future translational research effort may be applied in two directions. First, improvement of toxicity profile will extend the clinical benefit of ICL drugs. Reduced or reversible side effects will allow patients to tolerate prolonged treatment (104). Second, sensitization of cancer cells to ICL drugs could be another strategy to achieve additional tumor control. As the mechanisms of ICL repair emerge, key repair factors can be legitimate targets for sensitization. Although ICL-based chemotherapeutic drugs have been used since 70 years ago, much potential exists for future development and refinement of this classical regimen.
Conclusions
DNA ICLs is a complex and severely genotoxic lesion. Cellular mechanisms dealing with ICLs have proven to be complicated and await further investigation. On the one hand, failure in the proper repair of ICLs is likely a significant source of genomic instability and hence cancer development. The manifestations of FA, especially its cancer risk, have fully demonstrated this notion. On the other hand, the profound cytotoxicity of ICL-inducing agents yielded a major class of cancer chemotherapeutic drugs. This class of bifunctional alkylating drugs has continuously evolved with increasing variety and efficacy. Future studies of cellular mechanism of ICL response are expected to advance cancer etiology as well as therapy.
Acknowledgments
The authors thank Donald R Norwood (Department of Scientific Publication, The University of Texas MD Anderson Cancer Center) for his assistance in English editing.
Funding: This research is supported by the National Cancer Institute (CA127945 and CA097175 Project 3 to L.L.).
Footnote
Provenance and Peer Review: This article was commissioned by the Guest Editors (David J. Chen and Benjamin P.C. Chen) for the series “DNA Damage and Repair” published in Translational Cancer Research. The article has undergone external peer review.
Conflicts of Interest: Both authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.3978/j.issn.2218-676X.2013.03.01). The series “DNA Damage and Repair” was commissioned by the editorial office without any funding or sponsorship. The authors have no other conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- O'Donovan A, Davies AA, Moggs JG, et al. XPG endonuclease makes the 3' incision in human DNA nucleotide excision repair. Nature 1994;371:432-5. [PubMed]
- Goodman LS, Wintrobe MM. Nitrogen mustard therapy; use of methyl-bis (beta-chloroethyl) amine hydrochloride and tris (beta-chloroethyl) amine hydrochloride for Hodgkin's disease, lymphosarcoma, leukemia and certain allied and miscellaneous disorders. J Am Med Assoc 1946;132:126-32. [PubMed]
- McVey M. Strategies for DNA interstrand crosslink repair: insights from worms, flies, frogs, and slime molds. Environ Mol Mutagen 2010;51:646-58. [PubMed]
- Noll DM, Mason TM, Miller PS. Formation and repair of interstrand cross-links in DNA. Chem Rev 2006;106:277-301. [PubMed]
- Lawley PD, Phillips DH. DNA adducts from chemotherapeutic agents. Mutat Res 1996;355:13-40. [PubMed]
- Parrish JA, Fitzpatrick TB, Tanenbaum L, et al. Photochemotherapy of psoriasis with oral methoxsalen and longwave ultraviolet light. N Engl J Med 1974;291:1207-11. [PubMed]
- Garaycoechea JI, Crossan GP, Langevin F, et al. Genotoxic consequences of endogenous aldehydes on mouse haematopoietic stem cell function. Nature 2012;489:571-5. [PubMed]
- Langevin F, Crossan GP, Rosado IV, et al. Fancd2 counteracts the toxic effects of naturally produced aldehydes in mice. Nature 2011;475:53-8. [PubMed]
- Rosado IV, Langevin F, Crossan GP, et al. Formaldehyde catabolism is essential in cells deficient for the Fanconi anemia DNA-repair pathway. Nat Struct Mol Biol 2011;18:1432-4. [PubMed]
- Goodman LS, Wintrobe MM, Dameshek W, et al. Landmark article Sept. 21, 1946: Nitrogen mustard therapy. Use of methyl-bis(beta-chloroethyl)amine hydrochloride and tris(beta-chloroethyl)amine hydrochloride for Hodgkin’s disease, lymphosarcoma, leukemia and certain allied and miscellaneous disorders. By Louis S. Goodman, Maxwell M. Wintrobe, William Dameshek, Morton J. Goodman, Alfred Gilman and Margaret T. McLennan. JAMA 1984;251:2255-61. [PubMed]
- Swinyard EA, Goodman LS. Laboratory assay of anticonvulsant potency of some hydantoinates. Fed Proc 1946;5:205. [PubMed]
- Magaña-Schwencke N, Henriques JA, Chanet R, et al. The fate of 8-methoxypsoralen photoinduced crosslinks in nuclear and mitochondrial yeast DNA: comparison of wild-type and repair-deficient strains. Proc Natl Acad Sci U S A 1982;79:1722-6. [PubMed]
- Cole RS. Repair of interstrand cross-links in DNA induced by psoralen plus light. Yale J Biol Med 1973;46:492. [PubMed]
- Cole RS. Psoralen monoadducts and interstrand cross-links in DNA. Biochim Biophys Acta 1971;254:30-9. [PubMed]
- Cole RS, Levitan D, Sinden RR. Removal of psoralen interstrand cross-links from DNA of Escherichia coli: mechanism and genetic control. J Mol Biol 1976;103:39-59. [PubMed]
- Cheng S, Sancar A, Hearst JE. RecA-dependent incision of psoralen-crosslinked DNA by (A)BC excinuclease. Nucleic Acids Res 1991;19:657-63. [PubMed]
- Cheng S, Van Houten B, Gamper HB, et al. Use of psoralen-modified oligonucleotides to trap three-stranded RecA-DNA complexes and repair of these cross-linked complexes by ABC excinuclease. J Biol Chem 1988;263:15110-7. [PubMed]
- Sladek FM, Munn MM, Rupp WD, et al. In vitro repair of psoralen-DNA cross-links by RecA, UvrABC, and the 5'-exonuclease of DNA polymerase I. J Biol Chem 1989;264:6755-65. [PubMed]
- Miller RD, Prakash L, Prakash S. Genetic control of excision of Saccharomyces cerevisiae interstrand DNA cross-links induced by psoralen plus near-UV light. Mol Cell Biol 1982;2:939-48. [PubMed]
- Simon JA, Szankasi P, Nguyen DK, et al. Differential toxicities of anticancer agents among DNA repair and checkpoint mutants of Saccharomyces cerevisiae. Cancer Res 2000;60:328-33. [PubMed]
- Berardini M, Foster PL, Loechler EL. DNA polymerase II (polB) is involved in a new DNA repair pathway for DNA interstrand cross-links in Escherichia coli. J Bacteriol 1999;181:2878-82. [PubMed]
- Berardini M, Mackay W, Loechler EL. Evidence for a recombination-independent pathway for the repair of DNA interstrand cross-links based on a site-specific study with nitrogen mustard. Biochemistry 1997;36:3506-13. [PubMed]
- Räschle M, Knipscheer P, Enoiu M, et al. Mechanism of replication-coupled DNA interstrand crosslink repair. Cell 2008;134:969-80. [PubMed]
- Wang LC, Gautier J. The Fanconi anemia pathway and ICL repair: implications for cancer therapy. Crit Rev Biochem Mol Biol 2010;45:424-39. [PubMed]
- Liu T, Ghosal G, Yuan J, et al. FAN1 acts with FANCI-FANCD2 to promote DNA interstrand cross-link repair. Science 2010;329:693-6. [PubMed]
- Yamamoto KN, Kobayashi S, Tsuda M, et al. Involvement of SLX4 in interstrand cross-link repair is regulated by the Fanconi anemia pathway. Proc Natl Acad Sci U S A 2011;108:6492-6. [PubMed]
- Garcia-Higuera I, Taniguchi T, Ganesan S, et al. Interaction of the Fanconi anemia proteins and BRCA1 in a common pathway. Mol Cell 2001;7:249-62. [PubMed]
- Zheng H, Wang X, Warren AJ, et al. Nucleotide excision repair- and polymerase eta-mediated error-prone removal of mitomycin C interstrand cross-links. Mol Cell Biol 2003;23:754-61. [PubMed]
- Sarkar S, Davies AA, Ulrich HD, et al. DNA interstrand crosslink repair during G1 involves nucleotide excision repair and DNA polymerase zeta. EMBO J 2006;25:1285-94. [PubMed]
- Shen X, Jun S, O’Neal LE, et al. REV3 and REV1 play major roles in recombination-independent repair of DNA interstrand cross-links mediated by monoubiquitinated proliferating cell nuclear antigen (PCNA). J Biol Chem 2006;281:13869-72. [PubMed]
- Tucker MA, Coleman CN, Cox RS, et al. Risk of second cancers after treatment for Hodgkin’s disease. N Engl J Med 1988;318:76-81. [PubMed]
- Travis LB, Holowaty EJ, Bergfeldt K, et al. Risk of leukemia after platinum-based chemotherapy for ovarian cancer. N Engl J Med 1999;340:351-7. [PubMed]
- Wang W. Emergence of a DNA-damage response network consisting of Fanconi anaemia and BRCA proteins. Nat Rev Genet 2007;8:735-48. [PubMed]
- Stoepker C, Hain K, Schuster B, et al. SLX4, a coordinator of structure-specific endonucleases, is mutated in a new Fanconi anemia subtype. Nat Genet 2011;43:138-41. [PubMed]
- Vaz F, Hanenberg H, Schuster B, et al. Mutation of the RAD51C gene in a Fanconi anemia-like disorder. Nat Genet 2010;42:406-9. [PubMed]
- Lobitz S, Velleuer E. Guido Fanconi (1892-1979): a jack of all trades. Nat Rev Cancer 2006;6:893-8. [PubMed]
- Auerbach AD. A test for Fanconi’s anemia. Blood 1988;72:366-7. [PubMed]
- Joenje H, Patel KJ. The emerging genetic and molecular basis of Fanconi anaemia. Nat Rev Genet 2001;2:446-57. [PubMed]
- Meetei AR, de Winter JP, Medhurst AL, et al. A novel ubiquitin ligase is deficient in Fanconi anemia. Nat Genet 2003;35:165-70. [PubMed]
- Knipscheer P, Räschle M, Smogorzewska A, et al. The Fanconi anemia pathway promotes replication-dependent DNA interstrand cross-link repair. Science 2009;326:1698-701. [PubMed]
- Hoadley KA, Xue Y, Ling C, et al. Defining the molecular interface that connects the Fanconi anemia protein FANCM to the Bloom syndrome dissolvasome. Proc Natl Acad Sci U S A 2012;109:4437-42. [PubMed]
- Meetei AR, Medhurst AL, Ling C, et al. A human ortholog of archaeal DNA repair protein Hef is defective in Fanconi anemia complementation group M. Nat Genet 2005;37:958-63. [PubMed]
- Ciccia A, Ling C, Coulthard R, et al. Identification of FAAP24, a Fanconi anemia core complex protein that interacts with FANCM. Mol Cell 2007;25:331-43. [PubMed]
- Yan Z, Delannoy M, Ling C, et al. A histone-fold complex and FANCM form a conserved DNA-remodeling complex to maintain genome stability. Mol Cell 2010;37:865-78. [PubMed]
- Wang Y, Leung JW, Jiang Y, et al. FANCM and FAAP24 Maintain Genome Stability via Cooperative as Well as Unique Functions. Mol Cell 2013;49:997-1009. [PubMed]
- Gari K, Décaillet C, Delannoy M, et al. Remodeling of DNA replication structures by the branch point translocase FANCM. Proc Natl Acad Sci U S A 2008;105:16107-12. [PubMed]
- Gari K, Décaillet C, Stasiak AZ, et al. The Fanconi anemia protein FANCM can promote branch migration of Holliday junctions and replication forks. Mol Cell 2008;29:141-8. [PubMed]
- Collis SJ, Ciccia A, Deans AJ, et al. FANCM and FAAP24 function in ATR-mediated checkpoint signaling independently of the Fanconi anemia core complex. Mol Cell 2008;32:313-24. [PubMed]
- Huang M, Kim JM, Shiotani B, et al. The FANCM/FAAP24 complex is required for the DNA interstrand crosslink-induced checkpoint response. Mol Cell 2010;39:259-68. [PubMed]
- Schwab RA, Blackford AN, Niedzwiedz W. ATR activation and replication fork restart are defective in FANCM-deficient cells. EMBO J 2010;29:806-18. [PubMed]
- Shen X, Do H, Li Y, et al. Recruitment of fanconi anemia and breast cancer proteins to DNA damage sites is differentially governed by replication. Mol Cell 2009;35:716-23. [PubMed]
- Ohashi A, Zdzienicka MZ, Chen J, et al. Fanconi anemia complementation group D2 (FANCD2) functions independently of BRCA2- and RAD51-associated homologous recombination in response to DNA damage. J Biol Chem 2005;280:14877-83. [PubMed]
- Meindl A, Hellebrand H, Wiek C, et al. Germline mutations in breast and ovarian cancer pedigrees establish RAD51C as a human cancer susceptibility gene. Nat Genet 2010;42:410-4. [PubMed]
- Masson JY, Tarsounas MC, Stasiak AZ, et al. Identification and purification of two distinct complexes containing the five RAD51 paralogs. Genes Dev 2001;15:3296-307. [PubMed]
- Sung P, Krejci L, Van Komen S, et al. Rad51 recombinase and recombination mediators. J Biol Chem 2003;278:42729-32. [PubMed]
- Takata M, Sasaki MS, Tachiiri S, et al. Chromosome instability and defective recombinational repair in knockout mutants of the five Rad51 paralogs. Mol Cell Biol 2001;21:2858-66. [PubMed]
- Kim Y, Lach FP, Desetty R, et al. Mutations of the SLX4 gene in Fanconi anemia. Nat Genet 2011;43:142-6. [PubMed]
- Crossan GP, van der Weyden L, Rosado IV, et al. Disruption of mouse Slx4, a regulator of structure-specific nucleases, phenocopies Fanconi anemia. Nat Genet 2011;43:147-52. [PubMed]
- Fekairi S, Scaglione S, Chahwan C, et al. Human SLX4 is a Holliday junction resolvase subunit that binds multiple DNA repair/recombination endonucleases. Cell 2009;138:78-89. [PubMed]
- Muñoz IM, Hain K, Déclais AC, et al. Coordination of structure-specific nucleases by human SLX4/BTBD12 is required for DNA repair. Mol Cell 2009;35:116-27. [PubMed]
- Svendsen JM, Smogorzewska A, Sowa ME, et al. Mammalian BTBD12/SLX4 assembles a Holliday junction resolvase and is required for DNA repair. Cell 2009;138:63-77. [PubMed]
- Andersen SL, Bergstralh DT, Kohl KP, et al. Drosophila MUS312 and the vertebrate ortholog BTBD12 interact with DNA structure-specific endonucleases in DNA repair and recombination. Mol Cell 2009;35:128-35. [PubMed]
- Ciccia A, McDonald N, West SC. Structural and functional relationships of the XPF/MUS81 family of proteins. Annu Rev Biochem 2008;77:259-87. [PubMed]
- Niedernhofer LJ, Odijk H, Budzowska M, et al. The structure-specific endonuclease Ercc1-Xpf is required to resolve DNA interstrand cross-link-induced double-strand breaks. Mol Cell Biol 2004;24:5776-87. [PubMed]
- Kim Y, Spitz GS, Veturi U, et al. Regulation of multiple DNA repair pathways by the Fanconi anemia protein SLX4. Blood 2013;121:54-63. [PubMed]
- Wang AT, Sengerová B, Cattell E, et al. Human SNM1A and XPF-ERCC1 collaborate to initiate DNA interstrand cross-link repair. Genes Dev 2011;25:1859-70. [PubMed]
- Kratz K, Schöpf B, Kaden S, et al. Deficiency of FANCD2-associated nuclease KIAA1018/FAN1 sensitizes cells to interstrand crosslinking agents. Cell 2010;142:77-88. [PubMed]
- MacKay C, Déclais AC, Lundin C, et al. Identification of KIAA1018/FAN1, a DNA repair nuclease recruited to DNA damage by monoubiquitinated FANCD2. Cell 2010;142:65-76. [PubMed]
- Smogorzewska A, Desetty R, Saito TT, et al. A genetic screen identifies FAN1, a Fanconi anemia-associated nuclease necessary for DNA interstrand crosslink repair. Mol Cell 2010;39:36-47. [PubMed]
- Masuda Y, Ohmae M, Masuda K, et al. Structure and enzymatic properties of a stable complex of the human REV1 and REV7 proteins. J Biol Chem 2003;278:12356-60. [PubMed]
- Hlavin EM, Smeaton MB, Noronha AM, et al. Cross-link structure affects replication-independent DNA interstrand cross-link repair in mammalian cells. Biochemistry 2010;49:3977-88. [PubMed]
- Rall DP, Pechura CM. Effects on health of mustard gas. Nature 1993;366:398-9. [PubMed]
- Woutersen RA, Appelman LM, Van Garderen-Hoetmer A, et al. Inhalation toxicity of acetaldehyde in rats. III. Carcinogenicity study. Toxicology 1986;41:213-31. [PubMed]
- Brooks PJ, Theruvathu JA. DNA adducts from acetaldehyde: implications for alcohol-related carcinogenesis. Alcohol 2005;35:187-93. [PubMed]
- Baan R, Straif K, Grosse Y, et al. Carcinogenicity of alcoholic beverages. Lancet Oncol 2007;8:292-3. [PubMed]
- Butturini A, Gale RP, Verlander PC, et al. Hematologic abnormalities in Fanconi anemia: an International Fanconi Anemia Registry study. Blood 1994;84:1650-5. [PubMed]
- Rosenberg PS, Greene MH, Alter BP. Cancer incidence in persons with Fanconi anemia. Blood 2003;101:822-6. [PubMed]
- Hirsch B, Shimamura A, Moreau L, et al. Association of biallelic BRCA2/FANCD1 mutations with spontaneous chromosomal instability and solid tumors of childhood. Blood 2004;103:2554-9. [PubMed]
- Faivre L, Guardiola P, Lewis C, et al. Association of complementation group and mutation type with clinical outcome in fanconi anemia. European Fanconi Anemia Research Group. Blood 2000;96:4064-70. [PubMed]
- Reid S, Schindler D, Hanenberg H, et al. Biallelic mutations in PALB2 cause Fanconi anemia subtype FA-N and predispose to childhood cancer. Nat Genet 2007;39:162-4. [PubMed]
- Alter BP, Rosenberg PS, Brody LC. Clinical and molecular features associated with biallelic mutations in FANCD1/BRCA2. J Med Genet 2007;44:1-9. [PubMed]
- Wooster R, Neuhausen SL, Mangion J, et al. Localization of a breast cancer susceptibility gene, BRCA2, to chromosome 13q12-13. Science 1994;265:2088-90. [PubMed]
- Rahman N, Seal S, Thompson D, et al. PALB2, which encodes a BRCA2-interacting protein, is a breast cancer susceptibility gene. Nat Genet 2007;39:165-7. [PubMed]
- Cantor SB, Bell DW, Ganesan S, et al. BACH1, a novel helicase-like protein, interacts directly with BRCA1 and contributes to its DNA repair function. Cell 2001;105:149-60. [PubMed]
- Loveday C, Turnbull C, Ruark E, et al. Germline RAD51C mutations confer susceptibility to ovarian cancer. Nat Genet 2012;44:475-6; author reply 476. [PubMed]
- Nijnik A, Woodbine L, Marchetti C, et al. DNA repair is limiting for haematopoietic stem cells during ageing. Nature 2007;447:686-90. [PubMed]
- Lensch MW, Rathbun RK, Olson SB, et al. Selective pressure as an essential force in molecular evolution of myeloid leukemic clones: a view from the window of Fanconi anemia. Leukemia 1999;13:1784-9. [PubMed]
- Park JW, Pitot HC, Strati K, et al. Deficiencies in the Fanconi anemia DNA damage response pathway increase sensitivity to HPV-associated head and neck cancer. Cancer Res 2010;70:9959-68. [PubMed]
- Kutler DI, Wreesmann VB, Goberdhan A, et al. Human papillomavirus DNA and p53 polymorphisms in squamous cell carcinomas from Fanconi anemia patients. J Natl Cancer Inst 2003;95:1718-21. [PubMed]
- Shanafelt TD, Lin T, Geyer SM, et al. Pentostatin, cyclophosphamide, and rituximab regimen in older patients with chronic lymphocytic leukemia. Cancer 2007;109:2291-8. [PubMed]
- Mackey JR, Martin M, Pienkowski T, et al. Adjuvant docetaxel, doxorubicin, and cyclophosphamide in node-positive breast cancer: 10-year follow-up of the phase 3 randomised BCIRG 001 trial. Lancet Oncol 2013;14:72-80. [PubMed]
- Morley A, Stohlman F Jr. Cyclophosphamide-induced cyclical neutropenia. An animal model of a human periodic disease. N Engl J Med 1970;282:643-6. [PubMed]
- McElwain TJ, Powles RL. High-dose intravenous melphalan for plasma-cell leukaemia and myeloma. Lancet 1983;2:822-4. [PubMed]
- Barlogie B, Hall R, Zander A, et al. High-dose melphalan with autologous bone marrow transplantation for multiple myeloma. Blood 1986;67:1298-301. [PubMed]
- Facon T, Mary JY, Hulin C, et al. Melphalan and prednisone plus thalidomide versus melphalan and prednisone alone or reduced-intensity autologous stem cell transplantation in elderly patients with multiple myeloma (IFM 99-06): a randomised trial. Lancet 2007;370:1209-18. [PubMed]
- Rai KR, Peterson BL, Appelbaum FR, et al. Fludarabine compared with chlorambucil as primary therapy for chronic lymphocytic leukemia. N Engl J Med 2000;343:1750-7. [PubMed]
- Ajithkumar T, Parkinson C, Shamshad F, et al. Ifosfamide encephalopathy. Clin Oncol (R Coll Radiol) 2007;19:108-14. [PubMed]
- Einhorn LH. Treatment of testicular cancer: a new and improved model. J Clin Oncol 1990;8:1777-81. [PubMed]
- Querfeld C, Rosen ST, Kuzel TM, et al. Long-term follow-up of patients with early-stage cutaneous T-cell lymphoma who achieved complete remission with psoralen plus UV-A monotherapy. Arch Dermatol 2005;141:305-11. [PubMed]
- Stern RS. Carcinogenic risk of psoralen plus ultraviolet radiation therapy: evidence in humans. Natl Cancer Inst Monogr 1984;66:211-6. [PubMed]
- Gasparro FP. The role of PUVA in the treatment of psoriasis. Photobiology issues related to skin cancer incidence. Am J Clin Dermatol 2000;1:337-48. [PubMed]
- Cass I, Baldwin RL, Varkey T, et al. Improved survival in women with BRCA-associated ovarian carcinoma. Cancer 2003;97:2187-95. [PubMed]
- Boyd J, Sonoda Y, Federici MG, et al. Clinicopathologic features of BRCA-linked and sporadic ovarian cancer. JAMA 2000;283:2260-5. [PubMed]
- Abu-Surrah AS, Kettunen M. Platinum group antitumor chemistry: design and development of new anticancer drugs complementary to cisplatin. Curr Med Chem 2006;13:1337-57. [PubMed]


