
FAM83D promotes the proliferation and migration of hepatocellular carcinoma cells by inhibiting the FBXW7/MCL1 pathway
Introduction
Hepatocellular carcinoma (HCC), which accounts for >90% of primary tumors of the liver, is a highly prevalent tumor worldwide with rapidly increasing prevalence and mortality rates (1,2). The major risk factors of liver cirrhosis, including long-term virus infection, autoimmune hepatitis, and chronic alcohol use, are the common reasons for the development and progression of HCC. The morbidity and mortality of HCC remain high despite numerous interventions to prevent virus infection, reduce alcohol intake, and the like.
At present, surgery and interventional surgery are the main methods for treating HCC (3). However, the incidence of postoperative recurrence and metastasis is still high, and the 5-year survival rate (~12.5%) for patients with advanced HCC remains bleak (2). Additionally, while many researchers have studied the molecular mechanisms of HCC progression, due to its complexity, its specific pathogenesis has not yet been fully elucidated. Thus, it is crucial to further investigate the pathogenesis involved in the development and progression of HCC and thus develop promising and effective therapeutic strategies.
This research examined family with sequence similarity 83, member D (FAM83D), which is located on chromosome 20 encoding mitotic spindle-associated proteins (4). In lung cancer, FAM83D overexpression is associated with tumor size, lymph node metastasis, and TNM staging (5). FAM83D is also highly expressed in several cancers, such as ovarian cancer and metastatic lung adenocarcinoma (6,7). High FAM83D levels are associated with a poor clinical prognosis in colorectal cancer (8). However, the role and mechanism of FAM83D in the development of HCC have not been closely studied.
F-box and WD repeat domain containing 7 (FBXW7) is a member of the F-box family of proteins, and it is considered to be a tumor suppressor (9,10). FBXW7 targets various mammalian proteins, regulates cell proliferation, growth, and apoptosis, and promotes proteasome degradation. FBXW7 expression has been reported to be downregulated in patients with HCC (11), but the mechanism by which this occurs has not yet been closely investigated. A study has confirmed that the overexpression of FAM83D determines the alternative mechanism of FBXW7 inactivation in breast cancer (6). Similarly, FAM83D knockout can cause the degradation of neurogenic locus notch homolog protein 1 (Notch1) by causing the upregulation of FBXW7 (12,13). MCL1 is one of the FBXW7 substrates. FBXW7 interacts with MCL-1 and degrades through phosphorylation. FBXW7 can regulate the malignant potential of cholangiocarcinoma and cisplatin-induced apoptosis through MCL1. Thus, we hypothesized that FAM83D might affect the development and progression of HCC by downregulating the FBXW7 level and its related pathway.
There are now rich public repositories of ribonucleic acid (RNA)-sequencing and microarray data sets, such as The Cancer Genome Atlas (TCGA) (14) and the Gene Expression Omnibus (GEO) (15) databases. The current study sought to explore the effect of FAM83D on HCC development and progression. We found that the expression of FAM83D was significantly increased in tumor tissues and was associated with poor clinical prognosis in the TCGA-Liver Hepatocellular Carcinoma (TCGA-LIHC) data set. Then, we downregulated FAM83D expression in SNU449 and HUH7 cells transfected with FAM83D small interfering RNA (siRNA) and upregulated FAM83D expression by FMA83D overexpression transfection in BEL7402 cells. The ability of cell proliferation, migration, and apoptosis was evaluated. Additionally, FAM83D knockdown combined with FBXW7 siRNA or MCL1 overexpression transfection was performed to investigate the correlation between FAM83D and the FBXW7/MCL1 pathway. FBXW7 expression was found to be negatively correlated with both the FAM83D and MCL1 levels in TCGA-LIHC data set. Our study revealed a new molecular pathogenesis of HCC. Our findings highlight a novel mechanism for FAM83D promotes HCC cell migration and invasion through FBXW7/MCL1 signaling pathway. Therefore, it may be a potential new therapeutic target for HCC. We present the following article in accordance with the MDAR reporting checklist (available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-2069/rc).
Methods
Bioinformatics analysis
TCGA
TCGA (http://cancergenome.nih.gov/) database was used to obtain the FAM83D expression profile and clinicopathological characteristics of TCGA-LIHC patients. A Gene Expression Profiling Interactive Analysis (GEPIA) (http://gepia.cancer-pku.cn/index.html) was conducted to analyze and visualize the data from the tumor and non-tumor samples from TCGA database and the Genotype-Tissue Expression (GTEx) project. Pathological staging and related prognostic analyses were performed to examine differences in the transcription levels of FAM83D between tumor and non-tumor tissues. We used Kaplan-Meier curves for the prognostic analysis. This study was conducted in accordance with the Declaration of Helsinki (as revised in 2013) and the relevant policies in China.
GEO
Data sets with non-tumor tissues and HCC liver tumor tissues were obtained from the GEO database. The search keywords were “human”, “liver”, and “HCC”. Ultimately, GSE25097 and GSE76427 from the GEO repository were included in the analysis. R software was used to compare the expression levels of FAM83D between tumor tissues and non-tumor tissues.
Cell culture and transfection
Human HCC cell lines (i.e., 7721, SNU449, BEL7402, Hep3b, and HUH7) were purchased from Nanjing Cobioer Bioscienses Company. FAM83D siRNA (i.e., siFAM83D-1, siFAM83D-2, and siFAM83D-3), FBXW7 siRNA (siFBXW7), siRNA negative control (siNC), FAM83D overexpression plasmid pcDNA-FAM83D (pcDNA-FAM83D), MCL1 overexpression plasmid pcDNA-MCL1 (pcDNA-MCL1), and pcDNA negative control (pcDNA-NC) were purchased from Genechem Company (Shanghai, China). Lentivirus particles were used to infect the SNU449, HUH7 and BEL7402 cells. After the cells were transfected for 24 h, the cells were collected for further experiments.
RNA extraction, semi-quantitative and real-time PCR
We used TRIZOL reagent (Invitrogen) to extract total RNA from cell culture samples, and reverse transcriptions were performed. The amplification was achieved by using SYBR green master mix through LightCycler 480. The following primers were used: FAM83D, forward: 5'-GGGAAGGTTCACGAAAAGT-3', reverse, 5'-GGCCAGACAGAATTACCAA-3'; β-actin: forward: 5'-CATGTACGTTGCTATCCAGGC-3', reverse, 5'-CTCCTTAATGTCACGCACGCT-3'.
Cell proliferation assays
We used 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl-2H-tetrazolium bromide (MTT) assays (Yuanye Company) to assess cell proliferation. We cultured the SNU499, HUH7, and BEL7402 cells in 96-well culture plates of Roswell Park Memorial Institute Medium 1640 supplemented with 10% fetal bovine serum. When the cells were 50% confluent, we transfected the cells with the siRNAs or pcDNA. After transfection, 20 µL of MTS solution was added to each well and incubated for 4 h. The optical density value was measured at 492 nm with a microplate reader. We used an EDU-based DNA cell proliferation kit (Beyotime Company) to detect cell proliferation and performed all the experiments according to the manufacturers’ guidelines.
Wound healing assays
We cultured the HCC cells in 6-well plates. The cells were scratched with a 200-µL sterile pipette tip, washed 3 times with DPBS, and cultured for 24 h. Photographs were taken at 0 and 24 h after focusing on the same position.
Cell apoptosis
The transfected cells were stained with annexin V-PE and 7-AAD in accordance with the manufacturer’s protocol, resuspended in phosphate buffered solution at room temperature for 30 min, and analyzed by flow cytometry (BD Company). The percentage of apoptosis in the cells was detected by flow cytometry. The apoptosis-related proteins were then examined by western bot.
Western blot
The total protein of the cells was dissolved in RIPA lysis buffer (Servicebio Company) containing a 10% protease inhibitor cocktail. We used the standard bicinchoninic acid method to detect the protein concentration of the lysate supernatant. Next, we performed a western blot analysis. Anti-human FAM83D (1:1,000), anti-Bcl-2, anti-Bax, anti-PARP, anti-CAS3, anti-E-CAD, anti-N-CAD, anti-VIM, anti-FBXW7, anti-MCL1, and β-actin (1:5,000) primary antibodies were used. Finally, we used Image J software for the relative protein analysis.
Statistical analysis
We used SPSS 20.0 statistical software for the statistical test. The data were presented as the mean ± standard deviation (SD). Statistical analyses between two groups were performed using the Student’s t-test One-way analysis of variance (ANOVA) was used to determine the significance of differences between groups. A P value <0.05 (2-tailed) was considered statistically significant.
Results
FAM83D was upregulated in HCC tumor tissues
FAM83D has been confirmed to be significantly upregulated in multiple human cancer types. To better explore the role of FAM83D in HCC, we analyzed TCGA database and found that FAM83D was significantly overexpressed in TCGA-LIHC tissues compared to adjacent non-tumor tissues (see Figure 1A). We then further explored the GEO database to verify the FAM83D expression profile in HCC. As expected, we found that FAM83D was significantly more upregulated in HCC tumor tissues than adjacent tissues (see Figure 1B).
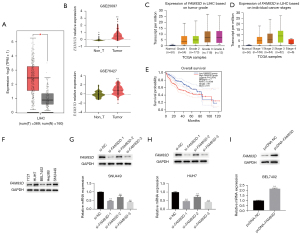
FAM83D was correlated with poor clinical outcomes
To investigate the effect of FAM83D elevation on clinical outcomes, we also evaluated the association between the expression of FAM83D and the clinical characteristics of TCGA-LIHC patients. A high level of FAM83D was significantly associated with a higher clinical stage and tumor grade (see Figure 1C,1D). The higher expression TCGA-LIHC group of FAM83D was more likely to have a higher grade and stage.
We then examined the relationship between FAM83D and HCC survival using the cBioPortal for Cancer Genomics. The Kaplan-Meier estimator was used to analyze the prognosis of TCGA-LIHC patients based on FAM83D expression. In Figure 1E, the blue line represents LIHC with relatively low expression of FAM83D, and the red line represents LIHC with high expression of FAM83D. We found that TCGA-LIHC patients with higher levels of FAM83D had significantly lower overall survival (pHR =0.0042, log-rank P=0.0046).
In short, our findings provide further evidence that FAM83D overexpression can be used as a poor prognosis indicator in HCC. Next, we detected FAM83D expression in the following 5 HCC cell lines: 721, SNU449, BEL7402, Hep3b, and HUH7.
FAM83D promoted HCC cell proliferation
The Western blot analysis showed that FAM83D expression was high in the SNU449 and HUH7 cells, and lowest in BEL7402 cells (see Figure 1F). We then downregulated the expression of FAM83D in the SNU449 and HUH7 cells transfected with FAM83D siRNA and upregulating its expression in the BEL7402 cells transfected with FMA83D overexpression (see Figure 1G-1I). We used 2 FAM83D siRNA sequences with efficient transfection for the subsequent in vitro studies, and the FAM83D protein expression level was significantly reduced in the SNU449 and HUH7 cells transfected with si-FAM83D-1 and si-FAM83D-2 (see Figure 1G,1H). Compared to the pcDNA-NC transfected cells, the FAM83D protein expression level was significantly increased after transfection with pcDNA-FAM83D in the BEL7402 cells (see Figure 1I).
To examine the role of FAM83D in the progression of HCC, we used MTT assays to evaluate the effects of FAM83D on the proliferation of HCC cell lines. Compared to the si-FAM83D-NC group, cell proliferation was significantly suppressed in the si-FAM83D-1 and si-FAM83D-2 groups. Conversely, cell proliferation was significantly enhanced following the overexpression of FAM83D (see Figure 2A).
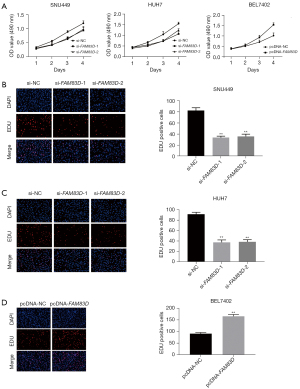
The EDU proliferation assays also showed that FAM83D knockdown strongly reduced cell proliferation (see Figure 2B,2C). We then tested the effect of FAM83D overexpression in the BEL7402 cells, and found that cell proliferation was significantly enhanced following the overexpression of FAM83D (see Figure 2D). Further, we observed that the colony formation ability of the si-FAM83D-1 and si-FAM83D-2 groups was significantly decreased compared to the FAM83D-NC group (see Figure 3A). Compared to the pc-NC group, the number of colonies of the BEL7402 cells transfected with pcDNA-FAM83D increased significantly (see Figure 3A). Taken together, these results indicated that FAM83D played an important role in accelerating HCC cell proliferation.
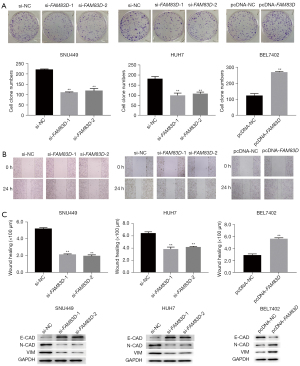
FAM83D regulated cell migration
It is well known that migration is a sign of tumor metastasis. We performed an in vitro scratch-healing experiment to explore the effect of FAM83D on cell migration. Consistent with our expectations, FAM83D knockdown reduced wound closure and uncoated migration through the Boyden chamber. We then evaluated the effect of FAM83D overexpression on cell migration in the HCC cells. Similarly, we found that the overexpression of FAM83D accelerated wound closure and uncoated migration through the Boyden chamber (see Figure 3B). Additionally, the western blotting analysis showed that the migration protein levels were significantly downregulated in the SNU449 and HUH7 cells following FAM83D knockdown. The migration protein levels were also significantly upregulated in the BEL7402 cells following FAM83D overexpression (see Figure 3C). These results indicated that FAM83D promoted cell migration in HCC.
FAM83D repressed apoptosis in HCC
We further explored the effect of FAM83D on the apoptosis of HCC cells. The annexin V-PE flow cytometry results showed that FAM83D knockdown promoted early cell apoptosis in the SNU449 and HUH7 cells. Additionally, the western blot analysis indicated that FAM83D knockdown significantly reduced the protein expression level of Bcl-2 but increased the protein expression of Bax and PARP (see Figure 4A,4B). Similarly, the flow cytometry analysis showed that high levels of FAM83D expression inhibited cell apoptosis (see Figure 4C). We also confirmed that the overexpression of FAM83D downregulated the expression of apoptotic-related proteins (see Figure 4C). Taken together, these results revealed that FAM83D repressed apoptosis, which might lead to the progression of HCC. Next, we examined the mechanisms of the role of FAM83D in the progression of HCC.
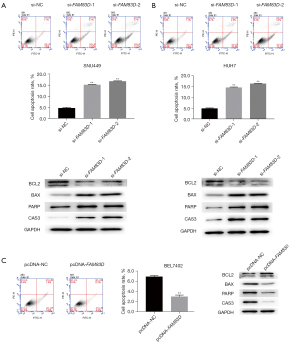
FAM83D enhanced in-vitro cell proliferation, migration and suppressed apoptosis by inhibiting the FBXW7/MCL1 pathway
Recently, a study showed that the overexpression of FAM83D determined the alternative mechanism of FBXW7 inactivation in breast cancer (6). Similarly, FAM83D knockout caused the degradation of Notch1 by causing the upregulation of FBXW7 (12). We thus hypothesized that the malignant behavior of HCC cells caused by the overexpression of FAM83D might be related to the downregulation of FBXW7.
First, western blots were used to examine the expression levels of FBXW7 and MCL1 in HCC cells. The results showed that FAM83D knockdown significantly increased FBXW7 expression but decreased the protein expression level of MCL1 (see Figure 5A). Conversely, the overexpression of FAM83D in BEL7402 had the opposite effect to that described above (see Figure 5A). These results suggested that FAM83D might promote HCC progression by affecting the FBXW7/MCL1 signaling pathway.
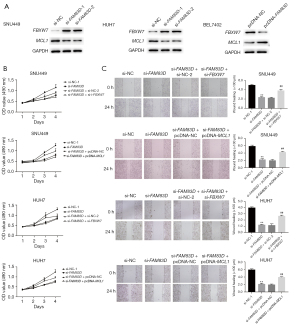
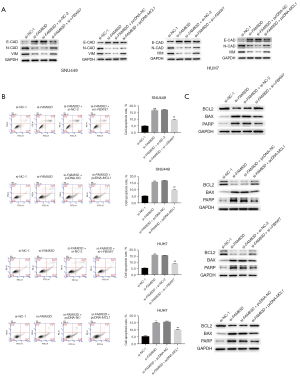
To further examine the effect of FAM83D on FBXW7/MCL1 signal transduction in HCC cells, the HCC cells were co-transfected with si-FAM83D and si-FBXW7 or with si-FAM83D and pc-MCL1. We found that either FBXW7 knockdown or MCL1 overexpression reduced the inhibitory effect of the downregulation of FAM83D on cell proliferation (see Figure 5B). Additionally, the scratch-healing experiment showed that the combination of FBXW7 knockdown or MCL1 overexpression reduced the inhibitory effect of FAM83D knockdown on cell migration (see Figure 5C). The Western blot analysis showed that both FBXW7 knockdown and MCL1 overexpression reversed the effect of reduced FAM83D expression on the migration protein level (see Figure 6A). Further, annexin V-PE flow cytometry and the western blot analysis showed that apoptosis was inhibited after the combined transfection with FBXW7 siRNA or MCL1 overexpression as compared to the only FAM83D siRNA group (see Figure 6B,6C). Thus, FBXW7 knockdown or MCL1 overexpression reduced the inhibitory effect of FAM83D knockdown on cell proliferation and migration, and the promotion of cell apoptosis. Our results indicated that FAM83D might regulate the proliferation and migration of HCC cells by inhibiting the FBXW7/MCL1 signaling pathway.
Discussion
HCC is characterized by a high incidence, a high degree of malignancy, and an extremely low survival rate. Surgical techniques and radiotherapy and chemotherapy are common treatment options for cancer patients, and there has been some progress in the targeted therapy of HCC; however, HCC continues to have a high risk of recurrence and a poor prognosis. In recent years, people have become more and more interested in identifying new genes that play a key role in the initiation and progression of cancer. An analysis of TCGA database identified a significantly increased level of FAM83D in HCC tissues, and revealed that FAM83D was closely related to the clinical prognosis of TCGA-LIHC patients. The FAM83D expression profile was then verified in the GEO database. In vitro studies showed that FAM83D facilitated the cell proliferation and migration and inhibited the cell apoptosis of HCC cells. We also found that FAM83D promoted the progression of HCC by inhibiting the FBXW7/MCL1 signaling pathway. These results suggested that FAM83D might be a promising target for HCC treatment.
FAM83D is located in the spindle and regulates the maintenance of the spindle, the process of mitosis, and cell division during mitosis (16,17). Studies have shown that the overexpression of FAM83D increases the migration and invasion of epithelial cells, which is related to a poor prognosis in breast cancer, lung cancer, and some other cancers (5,12). In our study, we demonstrated that FAM83D was an important molecule enhancing the proliferation and metastasis of HCC.
The analyses of TCGA and the GEO databases showed FAM83D was more upregulated in the HCC tumor tissues than the non-tumor tissues, and that a high level of FAM83D was closely related to a poor prognosis. Further, the results of the loss- and gain-of-function experiment in vitro showed that FAM83D promoted the proliferation and migration of HCC cells. The knockdown of FAM83D impaired tumor cell growth and migration, and promoted HCC cell apoptosis by increasing Bcl-2 and reducing Bax and PARP, while the upregulation of FAM83D reversed these results. Our findings provide novel insights into FAM83D-induced tumorigenesis and metastasis in HCC.
To achieve a deeper understanding of the role of FAM83D in the pathogenesis of HCC, we investigated the underlying pathways by which FAM83D affected HCC progression. F-box protein has received increasing attention due to its role in cancer. Among the F-box family, FBXW7 is thought to be a tumor suppressor involved in cell proliferation and migration in cancers, including lung, breast, gastric, and pancreatic cancer (10,18).
A study has shown that hepatocyte-specific FBXW7-deficient mice exhibit hepatomegaly and steatohepatitis (19). Further, research has shown that the expression level of FBXW7 affects the sensitivity of tumor cells to several chemotherapies (20,21). MCL1 is a key pro-survival member of the BCL2 protein family. MCL1 is one of the FBXW7 substrates. FBXW7 interacts with MCL-1 and degrades through phosphorylation (22). The loss of FBXW7 leads to the accumulation of MCL1, which leads to the chemotherapy resistance of cancer cells (23). Notably, it has been confirmed that the low expression of FBXW7 in squamous cell carcinoma increases the expression of MCL-1 and promotes resistance to standard chemotherapy (24). So we focused on the FBXW/MCL1 signaling pathway. In our study, TCGA-LIHC analysis showed that the expression of FAM83D was negatively correlated with FBXW7 expression, while FBXW7 expression level was negatively correlated with MCL1 expression level. we found that FAM83D enhanced the cell proliferation and migration and suppressed the cell apoptosis of the HCC cells by inhibiting the FBXW7/MCL1 signaling pathway. After the knockdown of FAM83D, the expression of FBXW7 protein was significantly increased and the expression of MCL1 protein was reduced. Additionally, we found that FBXW7 siRNA reversed the inhibitory effect of FAM83D knockdown on MCL1 protein expression, which clearly indicated that the oncogenic function of FAM83D is at least.
Acknowledgments
We would like to thank all those who participated in this study.
Funding: This work was supported by funding from
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-2069/rc
Data Sharing Statement: Available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-2069/dss
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-2069/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. This study was performed in accordance with the Declaration of Helsinki (as revised in 2013) and the relevant policies in China.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Sung H, Ferlay J, Siegel RL, et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin 2021;71:209-49. [Crossref] [PubMed]
- Wang W, Wei C. Advances in the early diagnosis of hepatocellular carcinoma. Genes Dis 2020;7:308-19. [Crossref] [PubMed]
- Greten TF, Lai CW, Li G, et al. Targeted and Immune-Based Therapies for Hepatocellular Carcinoma. Gastroenterology 2019;156:510-24. [Crossref] [PubMed]
- Meng T, Tong Z, Yang MY, et al. Immune implication of FAM83D gene in hepatocellular carcinoma. Bioengineered 2021;12:3578-92. [Crossref] [PubMed]
- Shi R, Sun J, Sun Q, et al. Upregulation of FAM83D promotes malignant phenotypes of lung adenocarcinoma by regulating cell cycle. Am J Cancer Res 2016;6:2587-98. [PubMed]
- Wang Z, Liu Y, Zhang P, et al. FAM83D promotes cell proliferation and motility by downregulating tumor suppressor gene FBXW7. Oncotarget 2013;4:2476-86. [Crossref] [PubMed]
- Ishii N, Araki K, Yokobori T, et al. Reduced FBXW7 expression in pancreatic cancer correlates with poor prognosis and chemotherapeutic resistance via accumulation of MCL1. Oncotarget 2017;8:112636-46. [Crossref] [PubMed]
- Hashimoto M, Kobayashi T, Tashiro H, et al. h-Prune is associated with poor prognosis and epithelial-mesenchymal transition in patients with colorectal liver metastases. Int J Cancer 2016;139:812-23. [Crossref] [PubMed]
- Davis RJ, Welcker M, Clurman BE. Tumor suppression by the Fbw7 ubiquitin ligase: mechanisms and opportunities. Cancer Cell 2014;26:455-64. [Crossref] [PubMed]
- Welcker M, Clurman BE. FBW7 ubiquitin ligase: a tumour suppressor at the crossroads of cell division, growth and differentiation. Nat Rev Cancer 2008;8:83-93. [Crossref] [PubMed]
- El-Mezayen H, Yamamura K, Yusa T, et al. MicroRNA-25 Exerts an Oncogenic Function by Regulating the Ubiquitin Ligase FBXW7 in Hepatocellular Carcinoma. Ann Surg Oncol 2021;28:7973-82. [Crossref] [PubMed]
- Mu Y, Zou H, Chen B, et al. FAM83D knockdown regulates proliferation, migration and invasion of colorectal cancer through inhibiting FBXW7/Notch-1 signalling pathway. Biomed Pharmacother 2017;90:548-54. [Crossref] [PubMed]
- Inuzuka H, Shaik S, Onoyama I, et al. SCF(FBW7) regulates cellular apoptosis by targeting MCL1 for ubiquitylation and destruction. Nature 2011;471:104-9. [Crossref] [PubMed]
- Barrett T, Wilhite SE, Ledoux P, et al. NCBI GEO: archive for functional genomics data sets--update. Nucleic Acids Res 2013;41:D991-5. [Crossref] [PubMed]
- Cancer Genome Atlas Research Network. The Cancer Genome Atlas Pan-Cancer analysis project. Nat Genet 2013;45:1113-20. [Crossref] [PubMed]
- Bozatzi P, Sapkota GP. The FAM83 family of proteins: from pseudo-PLDs to anchors for CK1 isoforms. Biochem Soc Trans 2018;46:761-71. [Crossref] [PubMed]
- Hua YQ, Zhang K, Sheng J, et al. FAM83D promotes tumorigenesis and gemcitabine resistance of pancreatic adenocarcinoma through the Wnt/β-catenin pathway. Life Sci 2021;287:119205. [Crossref] [PubMed]
- Forbes SA, Beare D, Boutselakis H, et al. COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res 2017;45:D777-83. [Crossref] [PubMed]
- Onoyama I, Suzuki A, Matsumoto A, et al. FBXW7 regulates lipid metabolism and cell fate decisions in the mouse liver. J Clin Invest 2011;121:342-54. [Crossref] [PubMed]
- Fang L, Yang Z, Zhou J, et al. Circadian Clock Gene CRY2 Degradation Is Involved in Chemoresistance of Colorectal Cancer. Mol Cancer Ther 2015;14:1476-87. [Crossref] [PubMed]
- Gstalder C, Liu D, Miao D, et al. Inactivation of FBXW7 Impairs dsRNA Sensing and Confers Resistance to PD-1 Blockade. Cancer Discov 2020;10:1296-311. [Crossref] [PubMed]
- Ren H, Koo J, Guan B, et al. The E3 ubiquitin ligases β-TrCP and FBXW7 cooperatively mediates GSK3-dependent Mcl-1 degradation induced by the Akt inhibitor API-1, resulting in apoptosis. Mol Cancer 2013;12:146. [Crossref] [PubMed]
- Wertz IE, Kusam S, Lam C, et al. Sensitivity to antitubulin chemotherapeutics is regulated by MCL1 and FBW7. Nature 2011;471:110-4. [Crossref] [PubMed]
- He L, Torres-Lockhart K, Forster N, et al. Mcl-1 and FBW7 control a dominant survival pathway underlying HDAC and Bcl-2 inhibitor synergy in squamous cell carcinoma. Cancer Discov 2013;3:324-37. [Crossref] [PubMed]
(English Language Editor: L. Huleatt)


