Magnetic resonance imaging-based radiomics analysis: predicting vascular invasion in breast invasive ductal carcinoma using different machine learning models
Highlight box
Key findings
• A combined model integrating magnetic resonance imaging (MRI) radiomics with clinical factors best predicted lymphovascular invasion (LVI) in breast invasive ductal carcinoma (IDC): area under the curve (AUC) =0.956 (training) and 0.778 (validation). Among eight machine learning models, extreme gradient boosting (XGBoost) performed best using only radiomics features (AUC =0.912 training, 0.706 validation). Independent LVI predictors: postmenopausal status, higher histological grade, sentinel lymph node metastasis, molecular subtype, and MRI maximum diameter ≥2 cm.
What is known and what is new?
• LVI is an adverse prognostic factor in breast cancer; conventional MRI has limited specificity and is subjective for preoperative LVI prediction.
• First systematic comparison of eight machine learning models (logistic regression, support vector machine, K‑nearest neighbors, random forest, ExtraTrees, XGBoost, LightGBM, multi‑layer perceptron) for MRI radiomics-based LVI prediction in IDC. The combined clinical‑radiomics model significantly outperforms pure radiomics or clinical models.
What is the implication, and what should change now?
• The combined model offers a non‑invasive preoperative tool to estimate LVI risk, supporting individualized treatment (e.g., neoadjuvant therapy or more aggressive surgery). Prospective multicenter validation and workflow standardization are needed before clinical use. Radiomics‑based LVI assessment may reduce unnecessary sentinel lymph node biopsies and guide adjuvant therapy de‑escalation or escalation.
Introduction
Globally, breast cancer has become the most common malignant tumor among women (1). The treatment of breast cancer has entered an era of individualized care centered on breast-conserving surgery, sentinel lymph node biopsy, and comprehensive systemic therapy. However, even with standardized treatment, some patients still face the risk of tumor recurrence and metastasis after surgery. Lymphovascular invasion (LVI) refers to the presence of tumor cells within defined endothelial spaces (e.g., lymphatic or blood vessels) in the peritumoral stroma of invasive breast cancer and/or the infiltration of tumor cells into peritumoral lymphatic or vascular spaces (2-4). Studies indicate that LVI is a recognized adverse prognostic factor for invasive breast cancer (5,6). The College of American Pathologists (CAP) considers LVI assessment the “gold standard” for cancer reporting. While core needle biopsies are essential for assessing the nature of neoplastic lesions, sampling limitations render them unsuitable for capturing architectural prognostic elements of tumor pathology, such as the critical presence of vascular invasion. Magnetic resonance imaging (MRI) offers higher sensitivity when mammography or ultrasound results are inconclusive, particularly for detecting multifocal disease or local recurrence (7). Recent international literature, including the St. Gallen Consensus, indicates a higher rate of local recurrence after breast-conserving surgery in LVI-positive breast cancer (8), though relevant studies remain scarce both domestically and internationally. Conventional MRI morphological assessment exhibits limitations such as high subjectivity and insufficient specificity in predicting LVI preoperatively (9). As an emerging technology, radiomics enables high-throughput extraction of quantitative features from medical images. Through a series of processes—image acquisition, segmentation, feature extraction and selection, model development, and evaluation—it aims to uncover tumor heterogeneity for more objective and accurate efficacy assessment. This study employs MRI radiomics to compare the predictive capabilities of various machine learning (ML) algorithms for detecting vascular invasion in invasive ductal carcinoma (IDC) of the breast. The objective is to establish and validate a combined predictive model integrating radiomics features with clinical-pathological factors, thereby providing novel imaging markers for non-invasive, preoperative assessment of LVI risk. This approach aims to guide personalized and precision clinical treatment decisions. We present this article in accordance with the TRIPOD reporting checklist (available at https://tcr.amegroups.com/article/view/10.21037/tcr-2025-1-2845/rc).
Methods
Patient population
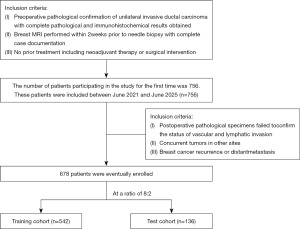
This study is a cross-sectional investigation conducted in compliance with the Declaration of Helsinki and its subsequent amendments and approved by the Ethics Committee of Guangdong Women and Children Hospital (No. KJ2024-254-01). Informed consent was not required due to the retrospective nature of the study. We retrospectively identified female patients diagnosed with pathologically confirmed IDC of the breast via MRI examination at our hospital between June 2021 and June 2025. Inclusion criteria: (I) preoperative pathological confirmation of unilateral IDC with complete pathological and immunohistochemical results; (II) breast MRI performed within 2 weeks prior to biopsy with complete case documentation; (III) no prior adjuvant therapy or surgical intervention. Exclusion criteria: (I) postoperative pathological examination failed to assess vascular/lymphatic invasion status; (II) concurrent tumors in other sites; (III) breast cancer recurrence or distant metastasis. A total of 678 patients aged 24–81 years (mean 50.31±10.44 years) were enrolled. Participants were randomly assigned to a training group (n=542) and a testing group (n=136) in an 8:2 ratio (see Figure 1).
Image acquisition
Using a Philips Ingenia 3.0 T MRI scanner from The Netherlands with a 16-channel phased array coil specifically designed for breast imaging. Primary scanning sequences and parameters: (I) transverse T1-weighted imaging (T1WI): repetition time (TR) 400–600 ms, echo time (TE) 8 ms, slice thickness 4 mm, slice spacing 0.4 mm, number of slices 40, matrix 212×240, field of view (FOV) 200 mm × 320 mm, number of echo planes (NEX) 1; (II) fat-suppressed transverse T2-weighted imaging (T2WI): TR 3,000–5,000 ms, TE 75 ms, slice thickness 4 mm, slice spacing 0.4 mm, number of slices 40, matrix 192×221, FOV 190 mm × 339 mm, NEX 1; (III) diffusion-weighted imaging (DWI) in the transverse plane: TR 6,550 ms, TE 99 ms, slice thickness 4 mm, slice spacing 0.4 mm, number of slices 40, matrix 212×142, FOV 400 mm × 340 mm, NEX 1, b-value 0, 800 mm2/s; (IV) transverse dynamic contrast-enhanced MRI (DCE-MRI): TR 4.4 ms, TE 2.2 ms, slice thickness 1.6 mm, slice spacing −0.8 mm, number of slices 187, matrix 328×392, FOV 280 mm × 339 mm, NEX 1. The contrast agent used was gadoteric acid glucamine injection (National Drug Approval Number H20153167, specification: 15 mL: 5.654 g). It was administered intravenously at a dose of 0.2 mmol/kg body weight with an infusion rate of 2.0 mL/s, followed by a 15 mL bolus of normal saline at the same rate. Dynamic contrast-enhanced scanning was performed post-injection, acquiring a total of 6 time phases.
Image analysis
Two radiologists with over 5 years of breast MRI diagnostic experience performed double-blind image interpretation. Disagreements were resolved through discussion to reach consensus. Breast MRI imaging features included: breast glandular tissue type [fibroglandular tissue (FGT)] (dense, non-dense); lesion type (mass, non-mass enhancement), time-signal intensity curve (TIC) pattern [influx (type I), plateau (type II), efflux (type III)], MRI axillary lymph node metastasis (present, absent), and MRI tumor maximum diameter (<2, ≥2 cm).
Determination of vascular invasion
Histopathological sections were stained with hematoxylin and eosin. Under light microscopy, clusters of cancer cells were observed within the spaces surrounded by flat endothelial cells, and these cancer cells were in continuity with the flat endothelial cells.
Clinical pathology and biological indicators
Clinical pathological characteristics primarily include age, age at menarche, menopausal status, family history of breast cancer, breast involvement, histological grade, sentinel lymph node status, and biological indicators. Biological markers primarily include estrogen receptor (ER), progesterone receptor (PR), human epidermal growth factor receptor 2 (HER2), and cell proliferation index (Ki-67). ER and PR assessment criteria: <1% positive nuclei are considered negative, ≥1% are positive (10). HER2 assessment criteria: scores of “−” or “+” are negative, “+++” is positive. Scores of “++” require further fluorescence in situ hybridization (FISH) testing; gene amplification indicates positive, no amplification indicates negative (11). Ki-67 assessment criteria: ≥14% is high expression, <14% is low expression. As outlined in the American Society of Clinical Oncology (ASCO)/CAP guidelines and the 2013 St. Gallen Consensus, breast cancer is classified based on immunohistochemistry (IHC) results into luminal A type (ER-positive and/or PR-positive, HER2-negative, Ki-67 <14%), luminal B (ER-positive and/or PR-positive), HER2-overexpressing (ER/PR-negative, HER2-positive), and triple-negative (ER/PR-negative, HER2-negative) subtypes (12,13).
Imageomics analysis
Image preprocessing: prior to feature extraction, all DCE-MRI images underwent the following preprocessing pipeline: N4 bias field correction, isotropic resampling to 1×1×1 mm3 voxel size, and gray-level discretization (quantization to 64 discrete bins) to standardize the data and reduce unwanted variance.
Image segmentation: the regions of interest (ROIs) were manually delineated. Using ITK-SNAP software (version 3.6.0, http://www.itk-snap.org), manually delineate the lesion ROI layer by layer on DCE-MRI phase-3 images. Initial delineation was performed by one attending physician, followed by verification by one associate chief physician. In cases of disagreement regarding tumor boundary delineation, a third associate chief physician was consulted to jointly determine the final ROI.
Feature extraction: high-dimensional radiomics features are extracted from each ROI, including geometric features, intensity features (first-order statistics), and texture features. Geometric features describe the three-dimensional shape of the tumor. Intensity features reflect the fundamental statistical distribution of voxel intensities within the tumor. Texture features are quantified based on the gray level co-occurrence matrix (GLCM), gray level run length matrix (GLRLM), gray level size zone matrix (GLSZM), gray level dependence matrix (GLDM), and neighborhood gray-tone difference matrix (NGTDM).
Feature selection: all features were initially screened using the Mann-Whitney U test (P<0.05). To eliminate highly redundant features, Spearman’s rank correlation coefficients were calculated. Any pair of features exhibiting a correlation coefficient exceeding 0.9 was deemed highly redundant. A greedy recursive removal strategy was then applied, retaining only the feature with higher discriminative power. This process ultimately yielded 23 features.
Feature modeling and label construction: further dimensionality reduction was achieved using least absolute shrinkage and selection operator (LASSO) regression (L1 regularization). The optimal regularization parameter λ was determined via 10-fold cross-validation, and features with non-zero coefficients were retained to construct the radiomics label (Rad-score). Employed logistic regression (LR), support vector machine (SVM), K-nearest neighbors (KNN), random forest, extreme random trees (ExtraTrees), extreme gradient boosting (XGBoost), light gradient boosting machine (LightGBM), and multi-layer perceptron (MLP) (XGBoost), LightGBM, and MLP. Five-fold cross-validation was employed during modeling to enhance generalization capability.
Clinical feature selection and cut-off point construction: predictive indicators (P<0.05) were screened from baseline clinical data, and a clinical model was constructed using the same ML workflow as for radiomics signatures. Radiomics signatures were further combined with significant clinical features to build a joint cut-off point model.
Model evaluation: the receiver operating characteristic (ROC) curve was used to assess the discriminatory ability of each model. Finally, decision curve analysis (DCA) was employed to evaluate the net benefit of the models at different clinical decision thresholds. The study workflow is illustrated in Figure 2.

Statistical analysis
Data analysis was performed using SPSS (version 26.0, IBM), R (version 3.6.3; https://www.r-project.org), and Python (version 3.5.6; http://www.python.org). For quantitative data, the Kolmogorov-Smirnov normality test was performed. If data followed a normal distribution, they were expressed as mean ± standard deviation, and comparisons between groups were conducted using the independent samples t-test (assuming equal variance). Otherwise, data were expressed as median (interquartile range), and comparisons between groups were performed using the Mann-Whitney U test. Count data were expressed as frequency (percentage). Intergroup comparisons employed either the χ2 test or Fisher’s exact test as appropriate. To validate model universality, beyond randomly allocating the entire sample to training and testing sets at an 8:2 ratio, we performed comparative analysis of baseline information across both sets to confirm randomness. For imageomics features in the training set, we first performed univariate screening using the Mann-Whitney U test. Features with P<0.05 advanced to subsequent rounds of screening, proceeding iteratively. Spearman correlation and LASSO regression were employed for feature dimensionality reduction and selection. ROC curves evaluated each model’s predictive capability for LVI in IDC of the breast, with area under the curve (AUC) and its 95% confidence interval (CI) calculated. P<0.05 indicated statistically significant results from hypothesis testing.
Results
Patient cohort and imaging characteristics
This study included 678 breast cancer patients, comprising 258 cases (38.05%) in the LVI-positive group and 420 cases (61.95%) in the LVI-negative group. All patients were randomly assigned in an 8:2 ratio to a training set (n=542) and a validation set (n=136). Baseline clinical and imaging characteristics for both groups are presented in Table 1. Within the training set, statistically significant differences (P<0.05) were observed between the LVI-positive and LVI-negative groups in age, menopausal status, histological grade, sentinel lymph node status, Ki-67, lesion type, MRI axillary lymph node status, and MR size; whereas in the validation set, significant intergroup differences were observed in histological grade, sentinel lymph node status, MRI axillary lymph node status, and MR size. Univariate and multivariate LR analyses revealed that menopausal status, histological grade, sentinel lymph node status, molecular subtype, and MRI size were independent predictors of LVI positivity in breast cancer patients (see Table 2). The clinical prediction model constructed based on multivariate analysis results indicated that LVI positivity was more likely to occur in postmenopausal patients [odds ratio (OR) =1.690; 95% CI: 1.174–2.433], with higher histological grade (OR =1.527; 95% CI: 1.107–2.107), and in cases with sentinel lymph node metastasis (OR =0.198; 95% CI: 0.137–0.285), molecular subtype (OR =0.740; 95% CI: 0.567–0.965), and MRI maximum diameter ≥2 cm (OR =2.059; 95% CI: 1.362–3.113).
Table 1
| Feature | Training cohort (n=542) | Testing cohort (n=136) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Training (n=542) | Negative (n=338) | Positive (n=204) | P value | Testing (n=136) | Negative (n=82) | Positive (n=54) | P value | ||
| Age (years) | 50.11±10.49 | 51.20±10.11 | 48.29±10.87 | 0.002* | 51.11±10.24 | 52.50±10.32 | 49.00±9.84 | 0.051 | |
| Age at menarche (years) | 12.23±0.89 | 12.26±0.98 | 12.18±0.73 | 0.81 | 12.18±0.70 | 12.16±0.68 | 12.22±0.74 | 0.45 | |
| Menopausal state | 0.009* | 0.30 | |||||||
| Premenopausal | 218 (40.22) | 151 (44.67) | 67 (32.84) | 64 (47.06) | 42 (51.22) | 22 (40.74) | |||
| Postmenopausal | 324 (59.78) | 187 (55.33) | 137 (67.16) | 72 (52.94) | 40 (48.78) | 32 (59.26) | |||
| Family history of breast cancer | 0.95 | 0.97 | |||||||
| No | 496 (91.51) | 310 (91.72) | 186 (91.18) | 122 (89.71) | 73 (89.02) | 49 (90.74) | |||
| Yes | 46 (8.49) | 28 (8.28) | 18 (8.82) | 14 (10.29) | 9 (10.98) | 5 (9.26) | |||
| Affecting the mammary glands | 0.57 | 0.21 | |||||||
| Left-sided | 278 (51.29) | 177 (52.37) | 101 (49.51) | 78 (57.35) | 43 (52.44) | 35 (64.81) | |||
| Right-sided | 264 (48.71) | 161 (47.63) | 103 (50.49) | 58 (42.65) | 39 (47.56) | 19 (35.19) | |||
| Histological grading | 0.002* | 0.03* | |||||||
| 1 | 69 (12.73) | 55 (16.27) | 14 (6.86) | 17 (12.50) | 14 (17.07) | 3 (5.56) | |||
| 2 | 328 (60.52) | 203 (60.06) | 125 (61.27) | 84 (61.76) | 44 (53.66) | 40 (74.07) | |||
| 3 | 145 (26.75) | 80 (23.67) | 65 (31.86) | 35 (25.74) | 24 (29.27) | 11 (20.37) | |||
| Sentinel lymph node | <0.001* | <0.001* | |||||||
| Positive | 193 (35.61) | 68 (20.12) | 125 (61.27) | 55 (40.44) | 20 (24.39) | 35 (64.81) | |||
| Negative | 349 (64.39) | 270 (79.88) | 79 (38.73) | 81 (59.56) | 62 (75.61) | 19 (35.19) | |||
| ER | 0.42 | 0.36 | |||||||
| Negative | 419 (77.31) | 257 (76.04) | 162 (79.41) | 104 (76.47) | 60 (73.17) | 44 (81.48) | |||
| Positive | 123 (22.69) | 81 (23.96) | 42 (20.59) | 32 (23.53) | 22 (26.83) | 10 (18.52) | |||
| PR | 0.23 | 0.42 | |||||||
| Negative | 370 (68.27) | 224 (66.27) | 146 (71.57) | 89 (65.44) | 51 (62.20) | 38 (70.37) | |||
| Positive | 172 (31.73) | 114 (33.73) | 58 (28.43) | 47 (34.56) | 31 (37.80) | 16 (29.63) | |||
| HER2 | 0.91 | 0.67 | |||||||
| Positive | 146 (26.94) | 90 (26.63) | 56 (27.45) | 29 (21.32) | 16 (19.51) | 13 (24.07) | |||
| Negative | 396 (73.06) | 248 (73.37) | 148 (72.55) | 107 (78.68) | 66 (80.49) | 41 (75.93) | |||
| Ki67 | 0.009* | >0.99 | |||||||
| High expression | 406 (74.91) | 240 (71.01) | 166 (81.37) | 105 (77.21) | 63 (76.83) | 42 (77.78) | |||
| Low expression | 136 (25.09) | 98 (28.99) | 38 (18.63) | 31 (22.79) | 19 (23.17) | 12 (22.22) | |||
| Molecular typing | 0.07 | 0.54 | |||||||
| Luminal A | 96 (17.71) | 68 (20.12) | 28 (13.73) | 22 (16.18) | 14 (17.07) | 8 (14.81) | |||
| Luminal B | 332 (61.25) | 194 (57.40) | 138 (67.65) | 82 (60.29) | 46 (56.10) | 36 (66.67) | |||
| HER2 overexpression | 53 (9.78) | 33 (9.76) | 20 (9.80) | 13 (9.56) | 8 (9.76) | 5 (9.26) | |||
| Triple-negative | 61 (11.25) | 43 (12.72) | 18 (8.82) | 19 (13.97) | 14 (17.07) | 5 (9.26) | |||
| Types of mammary gland tissue | 0.23 | 0.47 | |||||||
| Compact | 476 (87.82) | 292 (86.39) | 184 (90.20) | 116 (85.29) | 68 (82.93) | 48 (88.89) | |||
| Non-compact | 66 (12.18) | 46 (13.61) | 20 (9.80) | 20 (14.71) | 14 (17.07) | 6 (11.11) | |||
| Lesion type | 0.01* | 0.76 | |||||||
| Lump | 449 (82.84) | 291 (86.09) | 158 (77.45) | 113 (83.09) | 67 (81.71) | 46 (85.19) | |||
| Non-mass enhancement | 93 (17.16) | 47 (13.91) | 46 (22.55) | 23 (16.91) | 15 (18.29) | 8 (14.81) | |||
| TIC | 0.25 | 0.31 | |||||||
| I | 58 (10.70) | 34 (10.06) | 24 (11.76) | 14 (10.29) | 11 (13.41) | 3 (5.56) | |||
| II | 221 (40.77) | 147 (43.49) | 74 (36.27) | 55 (40.44) | 31 (37.80) | 24 (44.44) | |||
| III | 263 (48.52) | 157 (46.45) | 106 (51.96) | 67 (49.26) | 40 (48.78) | 27 (50.00) | |||
| MRI of axillary lymph nodes | 0.01* | <0.001* | |||||||
| Positive | 299 (55.17) | 172 (50.89) | 127 (62.25) | 73 (53.68) | 34 (41.46) | 39 (72.22) | |||
| Negative | 243 (44.83) | 166 (49.11) | 77 (37.75) | 63 (46.32) | 48 (58.54) | 15 (27.78) | |||
| MRI size | <0.001* | <0.001* | |||||||
| <2 cm | 180 (33.21) | 133 (39.35) | 47 (23.04) | 44 (32.35) | 40 (48.78) | 4 (7.41) | |||
| ≥2 cm | 362 (66.79) | 205 (60.65) | 157 (76.96) | 92 (67.65) | 42 (51.22) | 50 (92.59) | |||
Data are presented as mean ± standard deviation or n (%). *, P<0.05. ER, estrogen receptor; HER2, human epidermal growth factor receptor 2; MRI, magnetic resonance imaging; PR, progesterone receptor; TIC, time-signal intensity curve.
Table 2
| Variable | Univariate analysis | Multivariate analysis | |||
|---|---|---|---|---|---|
| OR (95% CI) | P value | OR (95% CI) | P value | ||
| Age | 0.291 (0.135, 0.447) | 0.66 | – | – | |
| Age at menarche | 0.061 (−0.094, 0.216) | 0.17 | – | – | |
| Menopausal state | 1.614 (1.172, 2.224) | 0.003* | 1.690 (1.174, 2.433) | 0.005* | |
| Family history of breast cancer | 1.013 (0.587, 1.748) | 0.96 | – | – | |
| Affecting the mammary glands | 0.987 (0.723, 1.346) | 0.93 | – | – | |
| Histological grading | NA | <0.001* | 1.527 (1.107, 2.107) | 0.01* | |
| Sentinel lymph node | 0.162 (0.115, 0.229) | <0.001* | 0.198 (0.137, 0.285) | <0.001* | |
| ER | 0.777 (0.533, 1.132) | 0.18 | – | – | |
| PR | 0.763 (0.545, 1.068) | 0.11 | – | – | |
| HER2 | 0.925 (0.650, 1.316) | 0.66 | – | – | |
| Ki-67 | 0.623 (0.428, 0.906) | 0.01* | 0.627 (0.377, 1.042) | 0.07 | |
| Molecular typing | NA | 0.03* | 0.740 (0.567, 0.965) | 0.02* | |
| Types of mammary gland tissue | 0.672 (0.412, 1.096) | 0.11 | – | – | |
| Lesion type | 1.528 (1.021, 2.288) | 0.03* | 1.348 (0.846, 2.150) | 0.20 | |
| TIC | NA | 0.47 | – | – | |
| MRI of axillary lymph nodes | 0.533 (0.388, 0.734) | <0.001* | 0.924 (0.636, 1.341) | 0.67 | |
| MRI size | 2.843 (1.978, 4.086) | <0.001* | 2.059 (1.362, 3.113) | <0.001* | |
*, P<0.05. CI, confidence interval; ER, estrogen receptor; HER2, human epidermal growth factor receptor 2; MRI, magnetic resonance imaging; NA, not applicable; OR, odds ratio; PR, progesterone receptor; TIC, time-signal intensity curve.
Radiomics feature models and performance evaluation
This study extracted a total of 1,197 radiomics features across six major categories from each patient’s imaging data. All features were extracted using the standardized workflow implemented by the PyRadiomics platform (http://pyradiomics.readthedocs.io). The specific number of features per category is detailed in Figure 3. Figure 3 illustrates the absolute count and the relative proportion (percentage) of the extracted features across the six different categories. Figure 4 displays the distribution of all features and their significance levels (P values) in comparisons between LVI groups. The feature correlation heatmap (Figure 5) indicates that morphological features such as tumor longest diameter, shortest diameter, and mean diameter exhibit the highest correlations, suggesting a degree of information overlap within the morphological feature set.



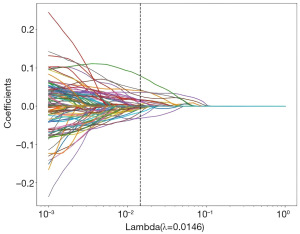
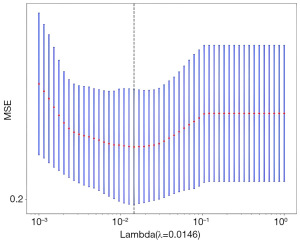
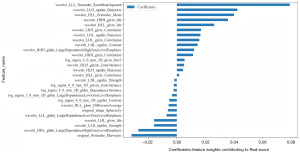
Lasso feature selection: select non-zero coefficients to establish a Rad-score LR model with the LASSO. The coefficients and mean squared error (MSE) from 10-fold cross-validation are shown in Figures 6,7, respectively. Figure 8 displays the coefficient values for the final selected non-zero features.
Model comparison: XGBoost demonstrated the best performance. Therefore, XGBoost was selected as the base model for constructing the Rad signature, as shown in Table 3.
Table 3
| Model | Task | AUC (95% CI) | Accuracy | Sensitivity | Specificity | PPV | NPV |
|---|---|---|---|---|---|---|---|
| LR | Train | 0.720 (0.6761–0.7635) | 0.679 | 0.662 | 0.689 | 0.562 | 0.772 |
| Test | 0.813 (0.7395–0.8875) | 0.765 | 0.796 | 0.744 | 0.672 | 0.847 | |
| SVM | Train | 0.830 (0.7945–0.8661) | 0.747 | 0.882 | 0.666 | 0.614 | 0.904 |
| Test | 0.776 (0.6932–0.8592) | 0.743 | 0.796 | 0.707 | 0.642 | 0.841 | |
| KNN | Train | 0.773 (0.7354–0.8110) | 0.734 | 0.559 | 0.840 | 0.679 | 0.759 |
| Test | 0.575 (0.4793–0.6702) | 0.581 | 0.463 | 0.659 | 0.472 | 0.651 | |
| Random forest | Train | 0.794 (0.7557–0.8318) | 0.736 | 0.681 | 0.769 | 0.641 | 0.800 |
| Test | 0.769 (0.6903–0.8476) | 0.713 | 0.759 | 0.683 | 0.612 | 0.812 | |
| ExtraTrees | Train | 0.684 (0.6390–0.7288) | 0.614 | 0.701 | 0.562 | 0.491 | 0.757 |
| Test | 0.752 (0.6715–0.8334) | 0.691 | 0.722 | 0.671 | 0.591 | 0.786 | |
| XGBoost | Train | 0.912 (0.8878–0.9370) | 0.832 | 0.863 | 0.814 | 0.736 | 0.908 |
| Test | 0.706 (0.6169–0.7946) | 0.662 | 0.741 | 0.610 | 0.556 | 0.781 | |
| LightGBM | Train | 0.834 (0.7992–0.8686) | 0.775 | 0.716 | 0.811 | 0.695 | 0.825 |
| Test | 0.755 (0.6736–0.8371) | 0.699 | 0.852 | 0.598 | 0.582 | 0.860 | |
| MLP | Train | 0.760 (0.7199–0.8004) | 0.716 | 0.652 | 0.754 | 0.616 | 0.782 |
| Test | 0.817 (0.7442–0.8904) | 0.743 | 0.833 | 0.683 | 0.634 | 0.862 |
AUC, area under the curve; CI, confidence interval; ExtraTrees, extreme random trees; GBM, gradient boosting machine; KNN, K-nearest neighbors; LR, logistic regression; MLP, multi-layer perceptron; NPV, negative prediction value; pCR, pathologic complete response; PPV, positive prediction value; SVM, support vector machine; XGBoost, extreme gradient boosting.
Rad signature was compared with LR, SVM, KNN, random forest, ExtraTrees, XGBoost, LightGBM, and MLP classifiers, with the radiological feature model ultimately yielding the optimal results. XGBoost achieved the best AUC value of 0.912 on the train set and 0.706 on the test set. The AUC values of each radiological feature model on the train and test sets are shown in Figure 9.

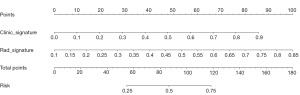
Nomgram
AUC: in the train cohort, both the clinic signature and Rad signature achieved perfect fit. In the test cohort, the clinic signature appeared overfitted, while the Rad signature maintained good fit. The nomogram combining the clinic signature and Rad signature using the LR algorithm yielded the best results, as shown in Table 4. Figure 10 displays the AUC values for the train and test sets. To compare the clinical signature, Rad signature, and nomogram, the DeLong test was employed (see Table 5).
Table 4
| Signature | Cohort | AUC (95% CI) | Accuracy | Sensitivity | Specificity | PPV | NPV |
|---|---|---|---|---|---|---|---|
| Clinic signature | Train | 0.889 (0.8626–0.9164) | 0.799 | 0.828 | 0.781 | 0.695 | 0.883 |
| Test | 0.727 (0.6392–0.8140) | 0.706 | 0.648 | 0.744 | 0.625 | 0.762 | |
| Rad signature | Train | 0.912 (0.8878–0.9370) | 0.832 | 0.863 | 0.814 | 0.736 | 0.908 |
| Test | 0.706 (0.6169–0.7946) | 0.662 | 0.741 | 0.610 | 0.556 | 0.781 | |
| Nomogram | Train | 0.956 (0.9394–0.9729) | 0.902 | 0.892 | 0.908 | 0.854 | 0.933 |
| Test | 0.778 (0.6997–0.8559) | 0.743 | 0.611 | 0.829 | 0.702 | 0.764 |
AUC, area under the curve; CI, confidence interval; NPV, negative prediction value; pCR, pathologic complete response; PPV, positive prediction value.

Table 5
| Cohort | Nomogram vs. clinic | Nomogram vs. rad |
|---|---|---|
| Train | 0.000399 | 0.066132 |
| Test | 0.010361 | 0.544511 |
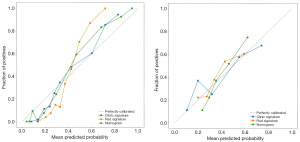
Calibration curve: the nomogram demonstrated good consistency between predicted and observed values in both the training and testing cohorts. The Hosmer-Lemeshow test P value indicated no statistically significant differences in clinical and radiographic characteristics between the training and testing cohorts, confirming that the nomogram achieved perfect fit in both groups (Table 6). Figure 11 shows the calibration curves for the training and testing cohorts.
Table 6
| Signature | P values |
|---|---|
| Clinic signature | 0.80 |
| 0.16 | |
| Rad signature | 0.64 |
| 0.55 | |
| Nomogram | 0.31 |
| 0.12 |
DCA: in this study, we also evaluated each model using DCA. The DCA for clinical features, radiographic features, and radiographic profiles is shown in Figure 12. In the test cohort, the DCA curves of the three models largely overlapped across a wide range of threshold probabilities. Figure 13 displays the nomogram for clinical use.

Discussion
This study established an imageomics model for predicting LVI in IDC based on preoperative breast DCE-MRI imageomics features, yielding the following key findings: the predictive model combining imageomics features with clinical-pathological indicators demonstrated optimal performance in predicting the presence or absence of LVI in IDC (train set AUC =0.956; test set AUC =0.778), demonstrating superior diagnostic performance compared to clinical-only or imaging-only models. This approach offers a promising adjunctive tool for non-invasive preoperative assessment of LVI risk.
LVI is recognized as an independent adverse prognostic factor for IDC of the breast and is associated with higher rates of lymph node involvement, local recurrence, and distant metastasis (2,14). Therefore, accurate preoperative assessment of LVI status is crucial for determining whether patients require axillary lymph node dissection and for planning postoperative chemoradiotherapy regimens (15). This study indicates that postmenopausal status, high nuclear grade, sentinel lymph node positivity, certain molecular subtypes (luminal B, HER2 positive, and triple-negative), and tumor longest diameter ≥2 cm on MRI are independent risk factors for LVI, consistent with most previous literature findings (16,17). Collectively, these factors reflect the aggressive biological behavior of the tumor. For instance, larger tumors or higher histological grades correlate with higher LVI detection rates. Studies by Lee et al. (18) and Rakha et al. (19) demonstrated that larger tumors exhibit higher LVI positivity rates. In Ugras’s study (20), the LVI positivity rate reached 23.6% in breast cancers with histological grades 2–3. This study shows that patients with grade 3 histology had an LVI positivity rate of 31.26%, consistent with literature values. This fully demonstrates that larger tumors and higher histological grades correlate with increased susceptibility to LVI involvement. Postmenopausal breast cancer often exhibits distinct molecular characteristics (e.g., higher HER2 expression, BRCA-related pathway activity) (21,22), which may partially explain the increased LVI risk in this population (23,24). The strong correlation between HER2 overexpression, high-proliferative subtypes, and LVI (25) further underscores the close relationship between intrinsic molecular drivers and the invasive phenotype of tumors. It is important to note that contemporary breast cancer management relies on a multifactorial decision-making framework, with primary weight given to age, hormone receptor status, HER2 expression, and genomic assays. LVI, while a robust prognosticator, typically functions as a complementary element within this framework. Our model aims to provide a preoperative estimate of this element, thereby offering additional contextual information that could assist in discussions regarding surgical planning and adjuvant therapy intensity, especially when other risk factors are equivocal.
Radiomics leverages high-throughput mining and analysis of large-scale quantitative features invisible to the naked eye within medical images, enabling precise characterization of tumor internal heterogeneity (26). This study extracted 1,197 features from DCE-MRI images, which underwent rigorous dimensionality reduction and filtering to generate a radiomics signature (Rad-score). Among univariate radiomics models, the XGBoost algorithm demonstrated the best performance (train set AUC =0.912, test set AUC =0.706). XGBoost, a gradient-boosted decision tree algorithm, excels at handling complex nonlinearities and feature interactions while preventing overfitting. This capability likely explains its superior performance on our high-dimensional feature data (27). However, the performance of pure radiomics models remains suboptimal and warrants further improvement.
Furthermore, the most noteworthy finding of this study is that the combined model achieved a predictive advantage where “1+1>2”, suggesting that clinical-pathological information (representing macroscopic biological status) and radiomics features (representing microscopic spatial heterogeneity) contain substantial complementary information. Constructing a logistic model using the aforementioned four multimodal indicators to predict LVI demonstrated optimal diagnostic performance. Its AUC value reached 0.956, surpassing any single indicator, with a sensitivity of 0.892 and specificity of 0.908. This indicates that clinical-pathological information and deeper radiomics features contain mutually exclusive predictive information. Clinical parameters reflect overall tumor biology (e.g., hormone receptor expression, Ki-67 levels), while radiomics parameters provide LVI-related information on tumor microstructural complexity, cellular density, and spatial heterogeneity of capillary formation. Integrating both enhances the accuracy of assessing tumor malignancy. Numerous radiomics studies have documented the advantages of integrating clinical and radiomics parameters (28,29).
Compared with recent similar studies, the joint model constructed in this study achieved comparable predictive performance. For example, Yang et al. (30) comprehensively collected patient imaging information and established a joint model based on MRI radiological features, radiomics features, and deep learning features. They found that the AUC for assessing breast cancer LVI positivity using the joint model reached 0.857. Liang et al. (31) developed four distinct ML models and deep learning models based on radiological features, clinical features, and radiomics features. The results showed that all five models achieved preoperative predictive efficacy exceeding 0.8 for invasive breast cancer LVI status. The aforementioned studies collectively demonstrate that utilizing a multi-feature approach based on MRI holds certain advantages for predicting LVI. However, the image sequences employed, feature extraction methods, and criteria for determining pathological outcomes varied across publications, making direct comparison of the established models challenging. There is an urgent need to standardize the methodologies and conduct external validation. While DCA suggested a potential for higher net clinical benefit with the combined model in the train set, its advantage was not pronounced in the independent test set, warranting further validation.
Limitations of this study: (I) this retrospective, single-center study carries inherent sample bias, and the generalizability of its model requires validation through multicenter prospective data; (II) ROIs were manually delineated and reviewed by two readers, yet time-consuming and inter-reader variability issues remain unavoidable. Future work may employ fully or semi-automated segmentation methods to enhance speed and consistency (32,33); (III) feature sources were limited to a single phase from DCE-MRI, lacking integration with multi-sequence information such as T2WI and DWI. Multi-parameter fusion has been demonstrated to provide additional predictive value (34); (IV) sample category imbalance existed, with disparities in the number of LVI-positive and LVI-negative cases. Future studies may employ more advanced algorithms for optimization.
Conclusions
In summary, this study demonstrates the feasibility of predicting vascular invasion in IDC using a combination of preoperative MRI radiomics and clinical features, with XGBoost showing advantages in modeling. The established combined nomogram model exhibits good discriminatory ability and clinical net benefit, offering potential as a non-invasive adjunct for preoperative risk assessment and personalized treatment strategy development. Future work may include conducting external validation on a large-scale cohort, establishing automated segmentation workflows, and further integrating additional MRI sequences or genomic information to develop higher-order comprehensive models.
Acknowledgments
None.
Footnote
Reporting Checklist: The authors have completed the TRIPOD reporting checklist. Available at https://tcr.amegroups.com/article/view/10.21037/tcr-2025-1-2845/rc
Data Sharing Statement: Available at https://tcr.amegroups.com/article/view/10.21037/tcr-2025-1-2845/dss
Peer Review File: Available at https://tcr.amegroups.com/article/view/10.21037/tcr-2025-1-2845/prf
Funding: None.
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://tcr.amegroups.com/article/view/10.21037/tcr-2025-1-2845/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki and its subsequent amendments. The study was approved by the Ethics Committee of Guangdong Women and Children Hospital (No. KJ2024-254-01). Informed consent was not required due to the retrospective nature of the study.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Siegel RL, Giaquinto AN, Jemal A. Cancer statistics, 2024. CA Cancer J Clin 2024;74:12-49. Erratum in: CA Cancer J Clin 2024;74:203.
- Kuhn E, Gambini D, Despini L, et al. Updates on Lymphovascular Invasion in Breast Cancer. Biomedicines 2023;11:968. [Crossref] [PubMed]
- Shen C, Lin C, Qu F, et al. Genomic spectra of lymphovascular invasion in breast cancer. Chin J Cancer Res 2025;37:138-53. [Crossref] [PubMed]
- Fan L, Wang J, Gao X, et al. Relationship between the ratio of mesenchymal cells/cancer cells and macrophages/cancer cells in vascular tumor thrombi and the clinical pathological characteristics and prognosis of patients with pT1-4N1-3M0 breast cancer. Chinese Journal of Breast Diseases 2020;14:23-31. (Electronic Edition).
- Yang Y, Wei H, Fu F, et al. Preoperative prediction of lymphovascular invasion of colorectal cancer by radiomics based on 18F-FDG PET-CT and clinical factors. Front Radiol 2023;3:1212382. [Crossref] [PubMed]
- Shen H, Zhang T, Wang S. A Prediction Model for Lymph Node Metastasis of Oral Squamous Cell Carcinoma Based on Multiple Risk Factors. Clin Exp Dent Res 2024;10:e70046. [Crossref] [PubMed]
- Sah AK, Choudhary RK, Sabrievna VA, et al. Male Breast Cancer: Epidemiology, Diagnosis, Molecular Mechanisms, Therapeutics, and Future Prospective. Oncol Res 2025;34:7. [Crossref] [PubMed]
- Zhong YM, Tong F, Shen J. Lympho-vascular invasion impacts the prognosis in breast-conserving surgery: a systematic review and meta-analysis. BMC Cancer 2022;22:102. [Crossref] [PubMed]
- Yang X, Wang X, Zuo Z, et al. Radiomics-based analysis of dynamic contrast-enhanced magnetic resonance image: A prediction nomogram for lymphovascular invasion in breast cancer. Magn Reson Imaging 2024;112:89-99. [Crossref] [PubMed]
- Allison KH, Hammond MEH, Dowsett M, et al. Estrogen and Progesterone Receptor Testing in Breast Cancer: American Society of Clinical Oncology/College of American Pathologists Guideline Update. Arch Pathol Lab Med 2020;144:545-63. [Crossref] [PubMed]
- Ahuja S, Khan AA, Zaheer S. Understanding the spectrum of HER2 status in breast cancer: From HER2-positive to ultra-low HER2. Pathol Res Pract 2024;262:155550. [Crossref] [PubMed]
- Wolff AC, Hammond MEH, Allison KH, et al. Human Epidermal Growth Factor Receptor 2 Testing in Breast Cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Focused Update. J Clin Oncol 2018;36:2105-22. [Crossref] [PubMed]
- Goldhirsch A, Winer EP, Coates AS, et al. Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol 2013;24:2206-23. [Crossref] [PubMed]
- Lin Y, Zhang Y, Fang H, et al. Survival and clinicopathological significance of blood vessel invasion in operable breast cancer: a systematic review and meta-analysis. Jpn J Clin Oncol 2023;53:35-45. [Crossref] [PubMed]
- Li Y, Yang J, Xiao P, et al. MRI Delta-Radiomics and Morphological Feature-Driven TabPFN Model for Preoperative Prediction of Lymphovascular Invasion in Invasive Breast Cancer. Technol Cancer Res Treat 2025;24:15330338251362050. [Crossref] [PubMed]
- Choi BB. Dynamic contrast enhanced-MRI and diffusion-weighted image as predictors of lymphovascular invasion in node-negative invasive breast cancer. World J Surg Oncol 2021;19:76. [Crossref] [PubMed]
- Wu X, Chen B, Li M, et al. Evaluation of Lymphovascular Invasion in Non-Specific Type Invasive Breast Cancer Patients with Mass Lesions Using Multimodal MRI Combined with Digital Breast Tomosynthesis. Journal of Practical Radiology 2025;41:599-602,613.
- Lee AH, Pinder SE, Macmillan RD, et al. Prognostic value of lymphovascular invasion in women with lymph node negative invasive breast carcinoma. Eur J Cancer 2006;42:357-62. [Crossref] [PubMed]
- Rakha EA, Martin S, Lee AH, et al. The prognostic significance of lymphovascular invasion in invasive breast carcinoma. Cancer 2012;118:3670-80. [Crossref] [PubMed]
- Ugras S, Stempel M, Patil S, et al. Estrogen receptor, progesterone receptor, and HER2 status predict lymphovascular invasion and lymph node involvement. Ann Surg Oncol 2014;21:3780-6. [Crossref] [PubMed]
- Anders CK, Hsu DS, Broadwater G, et al. Young age at diagnosis correlates with worse prognosis and defines a subset of breast cancers with shared patterns of gene expression. J Clin Oncol 2008;26:3324-30. [Crossref] [PubMed]
- Morrison DH, Rahardja D, King E, et al. Tumour biomarker expression relative to age and molecular subtypes of invasive breast cancer. Br J Cancer 2012;107:382-7. [Crossref] [PubMed]
- Shannon C, Smith IE. Breast cancer in adolescents and young women. Eur J Cancer 2003;39:2632-42. [Crossref] [PubMed]
- Guo S, Xie Y, Li Q, et al. Relationship between Dynamic Contrast-Enhanced MRI Features and Clinical-Pathological Characteristics of Invasive Breast Cancer and Vascular Invasion. Journal of PLA Medical Science 2025;50:847-54.
- Nishimura R, Osako T, Okumura Y, et al. An evaluation of lymphovascular invasion in relation to biology and prognosis according to subtypes in invasive breast cancer. Oncol Lett 2022;24:245. [Crossref] [PubMed]
- Qi YJ, Su GH, You C, et al. Radiomics in breast cancer: Current advances and future directions. Cell Rep Med 2024;5:101719. [Crossref] [PubMed]
- Chen Z, Wang Y, Ying MTC, et al. Interpretable machine learning model integrating clinical and elastosonographic features to detect renal fibrosis in Asian patients with chronic kidney disease. J Nephrol 2024;37:1027-39. [Crossref] [PubMed]
- Moran CJ. Editorial for "Intra- and Peritumoral Based Radiomics for Assessment of Lymphovascular Invasion in Invasive Breast Cancer". J Magn Reson Imaging 2024;59:626-7. [Crossref] [PubMed]
- Tenghui W, Xinyi L, Ziyi S, et al. Combination of ultrasound-based radiomics and deep learning with clinical data to predict response in breast cancer patients treated with neoadjuvant chemotherapy. Front Oncol 2025;15:1525285. [Crossref] [PubMed]
- Yang X, Fan X, Lin S, et al. Assessment of Lymphovascular Invasion in Breast Cancer Using a Combined MRI Morphological Features, Radiomics, and Deep Learning Approach Based on Dynamic Contrast-Enhanced MRI. J Magn Reson Imaging 2024;59:2238-49. [Crossref] [PubMed]
- Liang R, Li F, Yao J, et al. Predictive value of MRI-based deep learning model for lymphovascular invasion status in node-negative invasive breast cancer. Sci Rep 2024;14:16204. [Crossref] [PubMed]
- Li P, Ding J, Lim CS. VMDU-net: a dual encoder multi-scale fusion network for polyp segmentation with Vision Mamba and Cross-Shape Transformer integration. Front Artif Intell 2025;8:1557508. [Crossref] [PubMed]
- Wen C, Matsumoto M, Sawada M, et al. Seg2Link: an efficient and versatile solution for semi-automatic cell segmentation in 3D image stacks. Sci Rep 2023;13:7109. [Crossref] [PubMed]
- Xu J, Wang G, Wei Y, et al. Multi-parameter MRI deep learning model for lymphovascular invasion assessment in invasive breast ductal carcinoma: A multicenter, retrospective study. Clin Radiol 2025;88:107002. [Crossref] [PubMed]