
Prognostic significance of carboxypeptidase Q and its methylation in glioblastoma
Highlight box
Key findings
• Carboxypeptidase Q (CPQ) mRNA expression was higher in glioblastoma (GBM) tissues than in normal tissues.
• Low CPQ expression and high CPQ methylation were associated with better overall survival in GBM patients.
• Immune cells infiltration, immune markers and the tumor microenvironment were remarkably associated with CPQ expression in GBM.
What is known and what is new?
• GBM is the most common primary intracranial malignant tumor in adults, with poor prognosis.
• This study provided new insights into the prognostic role of CPQ and the potential role of CPQ in GBM tumor immunity.
What is the implication, and what should change now?
• CPQ is a promising biomarker for predicting prognosis in patients with GBM.
Introduction
Glioblastoma (GBM) is the most common primary intracranial malignant tumor in adults and is characterized by high aggressiveness and high mortality. The estimated annual incidence of GBM is 3.22 per 100,000 persons in the United States (1), and 5–8 per 100,000 persons in China (2). The five-year mortality rate of GBM in China is the third highest among all tumors, following pancreatic cancer and lung cancer. The current standard treatment includes maximum tumor resection, postoperative radiotherapy and concurrent temozolomide, followed by adjuvant chemotherapy (3). Despite the comprehensive treatment, GBM has a poor prognosis, with a median overall survival (OS) of 12 to 15 months, and a 5-year OS rate of only 6.8% (4,5). Almost all patients relapsed or progressed within a short period of time, with life expectancy dropping to 5–10 months after tumor recurrence (6). Since the Stupp regimen was first designed, treatment of GBM around the world has remained a bottleneck, with little improvement over the years (7). The complex biology of the disease, the high mutational heterogeneity observed between cancer cells, the presence of natural protection represented by the blood-brain barrier (BBB), and changes in the tumor microenvironment (TME) are all factors that explain the lack of improvement in the treatment of GBM (8-10).
Carboxypeptidase Q (CPQ), also known as plasma glutamic carboxypeptidase (PGCP), is an important member of the M28 family metal carboxypeptidase. CPQ is a secretory protein with five potential glycosylation sites (11), which locates in the vesicle compartment and is secreted into the extracellular space, mainly hydrolyzing circulating peptides in plasma. High levels of CPQ were found in both plasma (11) and thyroid (12). CPQ can be isolated from bovine spleen and rat liver (13), human placenta (11), human kidney (14) and thyroid (12). Literature reports and statistical analysis of The Cancer Genome Atlas (TCGA) database showed that CPQ was observably upregulated in about 60–70% of patients with cancer, including liver cancer, ovarian cancer, breast cancer, lung cancer and thyroid cancer (15). CPQ plays an important role in the hydrolysis of circulating peptides, and its activity depends on its dimerization (16), but the exact physiological substrates remain unclear. It helps to hydrolyze the neurotransmitter N-acetylaspartylglutamate (NAAG) in the brain to glutamate and N-acetylaspartate (NAA) (17), which indicates that it might play a role in glutamatergic synaptic dysfunction.
NAA is the second most abundant amino acid derivative in the brain, following glutamate (18). It contributes to neuronal mitochondrial metabolism, osmotic regulation of neuronal activity and axon-glial signaling (19). Possible connection was found to exist between NAA catabolism and oligodendrocyte progenitor cell (OPC) cell cycle arrest or oligodendrocyte differentiation (20). Long et al. showed that NAA and NAAG promoted the growth of glioma stem cells (GSC) and inhibited the differentiation of GSC. The increase of NAA or NAAG may enhance the phenotype of proliferative undifferentiated GSC (21). In GBM, CPQ might promote the growth of GSC and inhibit the differentiation of GSC by hydrolyzing NAAG into NAA, thus exacerbating the pathological changes of glioma. Here we attempt to analyze the relationship between CPQ expression and GBM.
Although GBM was refined into GBM, isocitrate dehydrogenase (IDH) wild-type in the 2021 WHO classification of tumors of the central nervous system (22), IDH mutation information in some databases was incomplete, so we mainly included traditional GBM patients for analysis. In this study, we investigated the differential expression of CPQ mRNA in GBM tissues and normal brain tissues, analyzed the association between CPQ expression and CPQ DNA methylation, and evaluated the prognostic significance of CPQ expression and DNA methylation in GBM. We then collected data from different databases and performed a multi-analysis of six public databases to assess the overall prognostic significance of CPQ. In addition, we assessed the potential connection between CPQ and GBM immune cell infiltration. Finally, the biological processes and signaling pathways involved in CPQ were examined by gene enrichment analysis to further analyze the mechanism of CPQ in GBM. We present this article in accordance with the REMARK reporting checklist (available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-2562/rc).
Methods
Database selection
First, we used GlioVis (http://gliovis.bioinfo.cnio.es/) to analyze the different expression of CPQ mRNA in GBM tissues and normal tissues based on the TCGA-GBM dataset (23). The University of ALabama at Birmingham CANcer data analysis Portal (UALCAN, http://ualcan.path.uab.edu/tutorial.html) was utilized to compare CPQ protein expression level between GBM tissues and normal tissues (24). Protein expression data were obtained from The National Cancer Institute’s Clinical Proteomic Tumor Analysis Consortium (CPTAC) (25).
Then we explored the correlation of CPQ mRNA expression and DNA methylation through cBioportal (https://www.cbioportal.org/) based on the TCGA-GBM dataset (26). We took advantage of MethSurv (https://biit.cs.ut.ee/methsurv/) to collect the DNA methylation data of CPQ in the TCGA-GBM dataset, then drew the heat map of methylation sites, and analyzed the prognosis of different methylation sites (27).
In addition, we collected data of GBM patients from TCGA, Chinese Glioma Genome Atlas (CGGA) and Gene Expression Omnibus (GEO) databases and verified the prognostic significance of CPQ using GlioVis (http://gliovis.bioinfo.cnio.es/). Inclusion criteria were GBM patients with complete clinical and transcriptional data. Parameters like mRNA and protein expression, methylation and sites, clinical characteristics, survival outcomes, immune cells infiltration, TME, immunoregulatory genes expression etc. were chosen and downloaded for different analyses. A total of 1,199 patients with pathologically confirmed GBM were included in this study (TCGA: 525 cases; CGGA: 220 cases; Gravendeel: 155 cases; LeeY: 191 cases; Murat: 80 cases; Nutt: 28 cases).
Biological function analysis
CAMOIP (https://www.camoip.net/) was used for Gene Ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis (28). GBM patients in the TCGA dataset were divided into high CPQ expression group and low CPQ expression group according to the median CPQ expression. The gene expression was considered different between the two groups, if the error detection rate was less than 0.05.
Correlation analysis of CPQ expression and immune cell infiltration
TIMER 2.0 (http://timer.cistrome.org/) was utilized to study the association of CPQ expression and immune cell infiltration (29). We made use of CAMOIP to evaluate the immune cell infiltration in TME. The population was divided into high CPQ group and low CPQ group according to the median CPQ expression value. The CIBERSORT algorithm was used to analyze the differences in the infiltration levels of 22 kinds of immune cells between the two groups, and the differences of tumor mutational burden (TMB), microsatellite instability (MSI) and Neoantigen scores between the two groups. The ESTIMATE algorithm was taken advantage to investigate the correlation between CPQ expression and the StromaScore and ImmuneScore. The relationship of CPQ expression and immunoregulatory factors expression was explored by Sangerbox 3.0 (http://vip.sangerbox.com/home.html).
Statistical analysis
R (version 4.1) and GraphPad Prism (version 8.0) were used to analyze the data. According to the median CPQ mRNA expression value in different datasets, CPQ was divided into two groups, namely, CPQ high group and CPQ low group. Similarly, CPQ higher methylated and lower methylated groups were established according to the median value of CPQ DNA methylation in the TCGA-GBM dataset. Analysis of the relationship between CPQ expression or its DNA methylation and a series of variables was performed using the Chi-squared test or Fisher’s exact test. The difference of the continuous variable with normal distribution between the two groups was determined by Student’s T-test, while the continuous variable with skewness distribution was determined by non-parametric test. The association between CPQ expression and CPQ DNA methylation level was identified by Pearson correlation coefficient. In addition, univariate and multivariate Cox regression models were used to investigate whether CPQ expression was an independent prognostic indicator for GBM patients. Kaplan-Meier curves were carried out to assess the prognostic significance of CPQ expression and CPQ DNA methylation. P values less than 0.05 on both sides were statistically significant.
Ethical statement
The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The Ethics Committee of the Fifth Affiliated Hospital of Sun Yat-sen University has confirmed that no ethical approval is required.
Results
Clinical and prognostic value of CPQ expression and its methylation
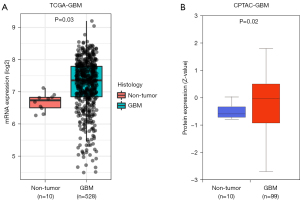
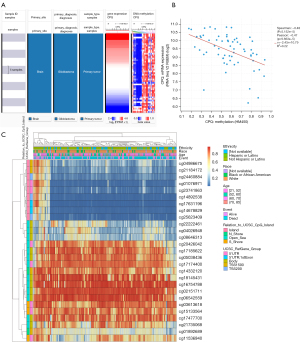
We collected and analyzed RNA expression data of 528 GBM tissues and 10 normal brain tissues from TCGA, and found that CPQ mRNA expression in GBM tissues was significantly higher than that in normal brain tissues (P=0.03, Figure 1A). The protein expression of CPQ in GBM tissue was also higher than that in normal brain tissue (P<0.05, Figure 1B). To clarify the cause of CPQ expression changes, we further investigated the genomic changes of CPQ, including gene mutation, copy number variation and DNA methylation. Results demonstrated that the DNA methylation level of CPQ was negatively correlated with its expression level (P<0.05; Figure 2A,2B). Figure 2C showed the heat map of methylation sites of CPQ. To further exclude other factors leading to abnormal CPQ expression, we also investigated the relationship between gene mutation or copy number variation and CPQ expression, and concluded that gene mutation and copy number variation were not related to CPQ expression (Figure S1).
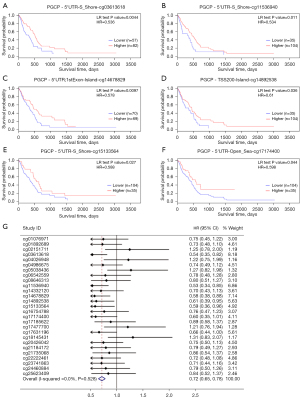
Then we explored the association between the methylation levels of CPQ at different methylation sites and the prognosis of GBM patients. Kaplan-Meier survival analysis displayed that higher methylation level at six CpG sites (CG03613618, CG14678829, CG11536940, CG14892538, CG15133564, and CG17174400) were associated with better OS (P<0.05, Figure 3A-3F) in GBM. Figure S2A-S2U showed methylation sites of CPQ that were irrelevant to OS. In a further meta-analysis of the correlation between all methylation sites and OS, we found that patients with higher CPQ methylation level had remarkably longer OS (Figure 3G), and higher CPQ methylation level was a protective factor.
Expression of CPQ in different subgroups of GBM
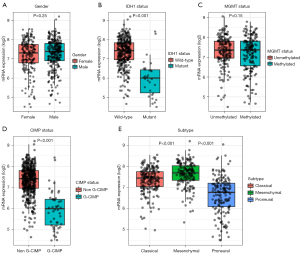
Non-parametric tests were applied to compare the differences of CPQ mRNA expression in subgroups classified by gender, IDH1 mutation, MGMT methylation, CIMP status and clinical subtypes (30) (Figure 4A-4E). CPQ mRNA expression in IDH1 wild-type was outstandingly higher than that in mutant type (P<0.001, Figure 4B). CPQ expression was extensively higher in the non-G-CIMP group than in the G-CIMP group (P<0.001, Figure 4D). The expression of CPQ in the Classical and Mesenchymal subtypes was expressively higher than that in proneural subtypes (P<0.001, Figure 4E). However, CPQ expression was not related to MGMT methylation or gender (Figure 4A,4C). Those results indicated that CPQ mRNA expression was closely related to a series of important clinical parameters.
Meta-analysis of the prognostic role of CPQ in GBM
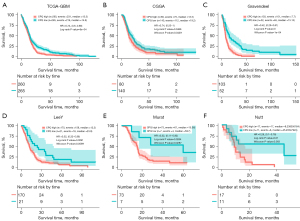
To verify the prognostic role of CPQ in GBM, we conducted validation in six datasets, including TCGA-GBM, CGGA, Gravendeel, LeeY, Murat, and Nutt (Figure 5A-5F). We found that in all six datasets, low CPQ expression was associated with better OS (P<0.05). Since the expression of CPQ was lower in IDH mutant population, we further narrowed the analysis population to GBM, IDH wildtype, based on TCGA-GBM, CGGA, and Gravendeel datasets in order to exclude the factor of IDH mutation causing the better prognosis in patients with low CPQ expression. Results demonstrated that CPQ was still significant in predicting the prognosis of IDH wildtype GBM patients in TCGA-GBM, and Gravendeel datasets (P<0.05), though not meaningful in the CGGA dataset (Figure S3A-S3C). Therefore, it can be concluded that low CPQ expression is a strong predictor of favorable OS in GBM patients, even in IDH wildtype GBM population.
Correlation analysis of CPQ mRNA expression and immune cell infiltration
To understand the functional role of CPQ in GBM, we identified potential biological processes and signaling pathways by GO and KEGG analysis. We compared the genome information of high and low CPQ expression subgroups in the TCGA-GBM dataset and found that the TOP20 biological processes relevant to the differentially expressed genes were almost all related to immunity (Figure 6A). KEGG analysis demonstrated that the differentially expressed genes were involved in several immune-related signaling pathways including cytokine-cytokine receptor interaction, chemokine signaling pathway, complement and coagulation cascades, Th17 cell differentiation, and intestinal immune network for IgA production (Figure 6B).
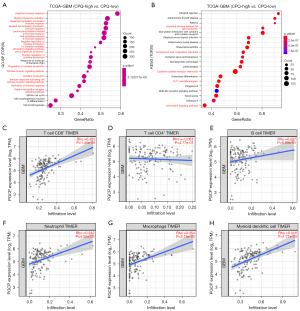
Subsequently, we investigated the association between CPQ expression and immune cell infiltration in GBM using TIMER 2.0 database, as shown in Figure 6C-6H, and found that CPQ mRNA expression was outstandingly correlated with CD8+ T cells, neutrophils, macrophages and DC cell infiltration, but not with CD4+ T cells and B cell infiltration. Then we analyzed the levels of immune cell infiltration between GBM samples with high and low CPQ expression in TCGA-GBM dataset through CIBERSORT. T cells regulatory, T cells gamma delta, NK cells resting, and Macrophages were found to be remarkably different in high and low CPQ expression specimens (Figure 7). Those findings demonstrated the close relationship between CPQ and tumor immunity.

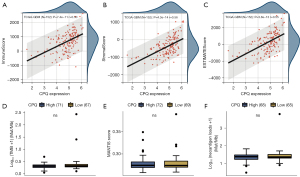
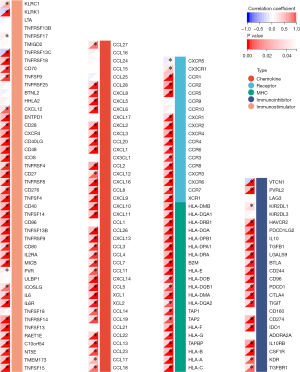
To further evaluate the association between CPQ and GBM tumor immunity, we performed a correlation analysis between CPQ expression and ESTIMATE score in the TCGA-GBM dataset, and the results showed that CPQ expression was meaningfully correlated with ImmuneScore, StromaScore, and the final ESTIMATE scores (Figure 8A-8C), suggesting that CPQ might affect the entire TME, including immune cells and stromal cells. However, there were no statistically significant differences in TMB (Figure 8D), MSI score (Figure 8E) and Neoantigen score (Figure 8F) between CPQ high and low samples. In addition, we explored the relation between expressions of CPQ and five types of immunoregulatory genes (chemokine, receptor, major histocompatibility complex (MHC), immunoinhibitor, immunostimulator) in TCGA-GBM dataset. Results revealed that CPQ expression was correlated with almost all immunomodulatory genes (Figure 9). It suggested that CPQ might affect TME by regulating immunomodulatory genes.
Discussion
In this study, the mRNA expression and the DNA methylation of CPQ in GBM, and their prognostic significance in clinic were analyzed based on TCGA database. We first found high CPQ mRNA expression in GBM tissues and a prominent negative correlation between CPQ mRNA expression and DNA methylation. We revealed that CPQ expression and methylation were closely related to a series of important characteristics of GBM, including clinical features and molecular types. Cox regression model confirmed that low CPQ expression and high CPQ methylation had good prognostic effects in GBM patients. Besides, six additional datasets analysis verified that low CPQ expression was associated with favorable OS. Finally, we demonstrated that immune cells infiltration, a series of immune markers and the TME were remarkably associated with CPQ expression in GBM. To the best of our knowledge, our study provided new insights into the prognostic role of CPQ and the potential role of CPQ in GBM tumor immunity.
CPQ had been reported to be significantly up-regulated in 60–70% of tumor patients, including liver cancer, ovarian cancer, breast cancer, lung cancer, thyroid cancer, etc. (15). However, the clinical, prognostic role of CPQ and its mechanism had never been studied in GBM. It had been found that CPQ helped hydrolyze the neurotransmitter NAAG in the brain into glutamate and NAA (17), which might promote GSC growth and inhibit glial differentiation (21). In this study, we systematically explored the relationship between CPQ expression and survival by obtaining data from public databases. Meta-analysis and multivariate Cox regression confirmed that low CPQ expression was correlated with longer OS in GBM patients, and low CPQ expression was an effective prognostic factor for OS. In a word, it suggested that CPQ is a promising biomarker for predicting prognosis in patients with GBM.
Our results presented that CPQ expression was markedly higher in IDH1 wild-type than in mutant type in GBM patients. Metabolomics analysis of gliomas showed that NAA and NAAG were significantly reduced in IDH1/2 mutant gliomas (31). Moreover, reaction rate and level of glutamate were lower in IDH mutant gliomas (31). Considering that CPQ can hydrolyze the NAAG into glutamate and NAA, the decreased level of NAA in IDH mutant cells may be related to the reduced catabolism of NAAG catalyzed by CPQ. However, NAAG levels also declined in IDH1/2 mutant cells, and the specific mechanism remains unclear. Hence, whether CPQ affects the occurrence and development of glioma by influencing the level of NAA has to be further explored.
There is increasing evidence that abnormal DNA methylation plays a crucial role in the progression of GBM (32). The CpG methylation of DNA was correlated with the molecular subtypes of GBM. In our study, we found a prominent negative connection between CPQ expression and CPQ DNA methylation. Six specific CpG sites in CPQ DNA promoters were identified, whose methylation were associated with better OS. Thus, the methylation of CPQ is a strong indicator of favorable OS and PFS. CpG Island methylation phenotype (CIMP) has become a distinct molecular subtype in many human malignancies, including GBM (33). In GBM, CIMP was associated with proneural subtype and IDH mutation status (33). IDH mutations were highly enriched in CIMP+ proneural subtype. In this study, the expression of CPQ decreased in IDH1 mutant GBM, and CPQ expression in the G-CIMP group was outstandingly lower than that in the non-G-CIMP group, suggesting that IDH1 might play an important role to connect CPQ and G-CIMP.
The interaction between GBM cells and other cells in the TME is an extremely complex phenomenon. The complexity of the GBM microenvironment influences the overall biology of GBM and enhances its resistance to treatment. GBM-associated macrophages/microglia cells accounted for 30% to 50% of GBM tumors (34). Those cells were considered to be one of the most critical cells in GBM TME, promoting glioma proliferation and migration, and secreting a variety of growth factors and cytokines (35). CD8+ T cytotoxic and CD4+ T helper cells are the main lymphocytes in GBM microenvironment (36). Our study revealed that CPQ expression was remarkably negatively correlated with immune cells infiltration including + T cells, neutrophils, macrophages and DC cells, and also associated with plenty of immune markers and the entire TME in GBM. Therefore, we speculated that CPQ could affect the prognosis of GBM patients by regulating the immune microenvironment. CPQ and tumor immunity deserve more exploration in the future.
According to the 2021 WHO classification of tumors of the central nervous system, the traditional GBM has been updated to GBM, IDH wild-type (22). Due to the limitation of data sources in the database, we could not obtain the information of IDH mutations in all patients, so we still included all GBM patients including IDH wild type and IDH mutation. Fortunately, there are three datasets (TCGA-GBM, CGGA, Gravendeel) containing information about IDH mutations in GBM patients, so we supplemented analysis of the survival significance of CPQ in GBM, IDH wild-type only patients in those datasets. That is a limitation of our study. The prognostic significance of CPQ and its methylation in GBM still needs to be verified in more GBM, IDH wild-type only cohorts.
Conclusions
In conclusion, we successfully screened and verified the significant correlation between CPQ and GBM prognosis. The low expression and high methylation of CPQ play a vital role in the occurrence, progression and prognosis of glioma, possibly working through regulation of immune microenvironment. However, the detailed mechanism needs to be further explored in biomedical experiments, which is also the limitation of our study. There have been no studies on the relationship between CPQ and GBM at present, but CPQ is a promising research target for the pathogenesis, diagnosis and treatment of GBM. Our research provided a basis for the further implementation of in vivo and in vitro experiments and a direction for GBM individualized treatment.
Acknowledgments
Funding: This work was supported by
Footnote
Reporting Checklist: The authors have completed the REMARK reporting checklist. Available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-2562/rc
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://tcr.amegroups.com/article/view/10.21037/tcr-22-2562/coif). The authors have no conflicts of interest to declare.
Ethical Statement:
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Ostrom QT, Price M, Neff C, et al. CBTRUS Statistical Report: Primary Brain and Other Central Nervous System Tumors Diagnosed in the United States in 2015-2019. Neuro Oncol 2022;24:v1-v95. [Crossref] [PubMed]
- Jiang T, Mao Y, Ma W, et al. CGCG clinical practice guidelines for the management of adult diffuse gliomas. Cancer Lett 2016;375:263-73. [Crossref] [PubMed]
- Stupp R, Taillibert S, Kanner A, et al. Effect of Tumor-Treating Fields Plus Maintenance Temozolomide vs Maintenance Temozolomide Alone on Survival in Patients With Glioblastoma: A Randomized Clinical Trial. JAMA 2017;318:2306-16. [Crossref] [PubMed]
- Wen PY, Weller M, Lee EQ, et al. Glioblastoma in adults: a Society for Neuro-Oncology (SNO) and European Society of Neuro-Oncology (EANO) consensus review on current management and future directions. Neuro Oncol 2020;22:1073-113. [Crossref] [PubMed]
- Ostrom QT, Cioffi G, Gittleman H, et al. CBTRUS Statistical Report: Primary Brain and Other Central Nervous System Tumors Diagnosed in the United States in 2012-2016. Neuro Oncol 2019;21:v1-v100. [Crossref] [PubMed]
- Wu W, Lamborn KR, Buckner JC, et al. Joint NCCTG and NABTC prognostic factors analysis for high-grade recurrent glioma. Neuro Oncol 2010;12:164-72. [Crossref] [PubMed]
- Tan AC, Ashley DM, López GY, et al. Management of glioblastoma: State of the art and future directions. CA Cancer J Clin 2020;70:299-312. [Crossref] [PubMed]
- Nunno VD, Franceschi E, Gatto L, et al. BET inhibitors: the promise of a new generation of immunotherapy in glioblastoma. Immunotherapy 2021; Epub ahead of print. [Crossref] [PubMed]
- Di Nunno V, Franceschi E, Tosoni A, et al. Glioblastoma: Emerging Treatments and Novel Trial Designs. Cancers (Basel) 2021;13:3750. [Crossref] [PubMed]
- Di Nunno V, Franceschi E, Tosoni A, et al. Treatment of recurrent glioblastoma: state-of-the-art and future perspectives. Expert Rev Anticancer Ther 2020;20:785-95. [Crossref] [PubMed]
- Gingras R, Richard C, El-Alfy M, et al. Purification, cDNA cloning, and expression of a new human blood plasma glutamate carboxypeptidase homologous to N-acetyl-aspartyl-alpha-glutamate carboxypeptidase/prostate-specific membrane antigen. J Biol Chem 1999;274:11742-50. [Crossref] [PubMed]
- Dunn AD, Myers HE, Dunn JT. The combined action of two thyroidal proteases releases T4 from the dominant hormone-forming site of thyroglobulin. Endocrinology 1996;137:3279-85. [Crossref] [PubMed]
- McDonald JK, Zeitman BB, Ellis S. Detection of a lysosomal carboxypeptidase and a lysosomal dipeptidase in highly-purified dipeptidyl aminopeptidase I (cathepsin C) and the elimination of their activities from preparations used to sequence peptides. Biochem Biophys Res Commun 1972;46:62-70. [Crossref] [PubMed]
- Dolenc I, Mihelic M. Purification and primary structure determination of human lysosomal dipeptidase. Biol Chem 2003;384:317-20. [Crossref] [PubMed]
- Zajc T, Suban D, Rajković J, et al. Baculoviral expression and characterization of human recombinant PGCP in the form of an active mature dimer and an inactive precursor protein. Protein Expr Purif 2011;75:119-26. [Crossref] [PubMed]
- Dolenc I, Pain R, Turk V. Presence of the propeptide on recombinant lysosomal dipeptidase controls both activation and dimerization. Biol Chem 2007;388:47-51. [Crossref] [PubMed]
- Pruitt KD, Tatusova T, Brown GR, et al. NCBI Reference Sequences (RefSeq): current status, new features and genome annotation policy. Nucleic Acids Res 2012;40:D130-5. [Crossref] [PubMed]
- Mehta V, Namboodiri MA. N-acetylaspartate as an acetyl source in the nervous system. Brain Res Mol Brain Res 1995;31:151-7. [Crossref] [PubMed]
- Moffett JR, Ross B, Arun P, et al. N-Acetylaspartate in the CNS: from neurodiagnostics to neurobiology. Prog Neurobiol 2007;81:89-131. [Crossref] [PubMed]
- Mattan NS, Ghiani CA, Lloyd M, et al. Aspartoacylase deficiency affects early postnatal development of oligodendrocytes and myelination. Neurobiol Dis 2010;40:432-43. [Crossref] [PubMed]
- Long PM, Moffett JR, Namboodiri AMA, et al. N-acetylaspartate (NAA) and N-acetylaspartylglutamate (NAAG) promote growth and inhibit differentiation of glioma stem-like cells. J Biol Chem 2013;288:26188-200. [Crossref] [PubMed]
- Louis DN, Perry A, Wesseling P, et al. The 2021 WHO Classification of Tumors of the Central Nervous System: a summary. Neuro Oncol 2021;23:1231-51. [Crossref] [PubMed]
- Bowman RL, Wang Q, Carro A, et al. GlioVis data portal for visualization and analysis of brain tumor expression datasets. Neuro Oncol 2017;19:139-41. [Crossref] [PubMed]
- Chandrashekar DS, Bashel B, Balasubramanya SAH, et al. UALCAN: A Portal for Facilitating Tumor Subgroup Gene Expression and Survival Analyses. Neoplasia 2017;19:649-58. [Crossref] [PubMed]
- Wu P, Heins ZJ, Muller JT, et al. Integration and Analysis of CPTAC Proteomics Data in the Context of Cancer Genomics in the cBioPortal. Mol Cell Proteomics 2019;18:1893-8. [Crossref] [PubMed]
- Gao J, Aksoy BA, Dogrusoz U, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 2013;6:pl1. [Crossref] [PubMed]
- Modhukur V, Iljasenko T, Metsalu T, et al. MethSurv: a web tool to perform multivariable survival analysis using DNA methylation data. Epigenomics 2018;10:277-88. [Crossref] [PubMed]
- Lin A, Qi C, Wei T, et al. CAMOIP: a web server for comprehensive analysis on multi-omics of immunotherapy in pan-cancer. Brief Bioinform 2022;23:bbac129. [Crossref] [PubMed]
- Li T, Fu J, Zeng Z, et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res 2020;48:W509-14. [Crossref] [PubMed]
- Wang Q, Hu B, Hu X, et al. Tumor Evolution of Glioma-Intrinsic Gene Expression Subtypes Associates with Immunological Changes in the Microenvironment. Cancer Cell 2017;32:42-56.e6. [Crossref] [PubMed]
- Reitman ZJ, Jin G, Karoly ED, et al. Profiling the effects of isocitrate dehydrogenase 1 and 2 mutations on the cellular metabolome. Proc Natl Acad Sci U S A 2011;108:3270-5. [Crossref] [PubMed]
- Kraboth Z, Galik B, Tompa M, et al. DNA CpG methylation in sequential glioblastoma specimens. J Cancer Res Clin Oncol 2020;146:2885-96. [Crossref] [PubMed]
- Noushmehr H, Weisenberger DJ, Diefes K, et al. Identification of a CpG island methylator phenotype that defines a distinct subgroup of glioma. Cancer Cell 2010;17:510-22. [Crossref] [PubMed]
- Rossi ML, Hughes JT, Esiri MM, et al. Immunohistological study of mononuclear cell infiltrate in malignant gliomas. Acta Neuropathol 1987;74:269-77. [Crossref] [PubMed]
- Brandenburg S, Müller A, Turkowski K, et al. Resident microglia rather than peripheral macrophages promote vascularization in brain tumors and are source of alternative pro-angiogenic factors. Acta Neuropathol 2016;131:365-78. [Crossref] [PubMed]
- Tamma R, Ingravallo G, Annese T, et al. Tumor Microenvironment and Microvascular Density in Human Glioblastoma. Cells 2022;12:11. [Crossref] [PubMed]


